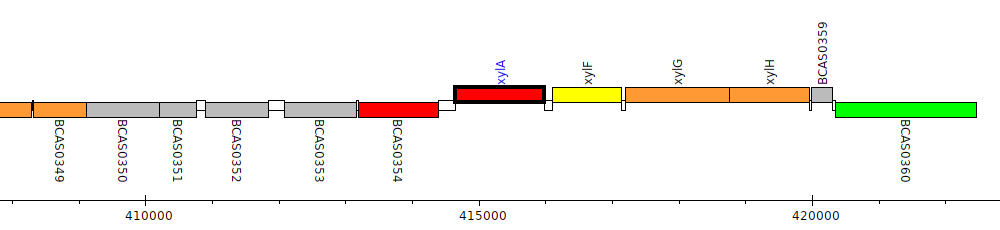
Burkholderia cenocepacia J2315, BCAS0355 (xylA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0009045 | xylose isomerase activity | Inferred from Experiment | |
||
| Biological Process | GO:0042732 | D-xylose metabolic process | Inferred from Experiment | |
||
| Biological Process | GO:0005975 | carbohydrate metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PS51415
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0009045 | xylose isomerase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS51415
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG (InterPro) | 00040 | Pentose and glucuronate interconversions | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG (InterPro) | 00051 | Fructose and mannose metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj00051 | Fructose and mannose metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00040 | Pentose and glucuronate interconversions | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00688 | Xylose isomerase signature | IPR001998 | Xylose isomerase | 206 | 225 | 9.6E-86 |
| SUPERFAMILY | SSF51658 | IPR036237 | Xylose isomerase-like superfamily | 3 | 438 | 2.35E-166 | |
| PRINTS | PR00688 | Xylose isomerase signature | IPR001998 | Xylose isomerase | 332 | 343 | 9.6E-86 |
| Gene3D | G3DSA:3.20.20.150 | 2 | 439 | 1.7E-175 | |||
| PRINTS | PR00688 | Xylose isomerase signature | IPR001998 | Xylose isomerase | 228 | 252 | 9.6E-86 |
| TIGRFAM | TIGR02630 | xylose_isom_A: xylose isomerase | IPR013452 | Xylose isomerase, bacterial-type | 3 | 436 | 1.2E-201 |
| PRINTS | PR00688 | Xylose isomerase signature | IPR001998 | Xylose isomerase | 91 | 113 | 9.6E-86 |
| PRINTS | PR00688 | Xylose isomerase signature | IPR001998 | Xylose isomerase | 299 | 310 | 9.6E-86 |
| PRINTS | PR00688 | Xylose isomerase signature | IPR001998 | Xylose isomerase | 179 | 200 | 9.6E-86 |
| PRINTS | PR00688 | Xylose isomerase signature | IPR001998 | Xylose isomerase | 270 | 297 | 9.6E-86 |
| ProSiteProfiles | PS51415 | Xylose isomerase family profile. | IPR001998 | Xylose isomerase | 34 | 436 | 186.891 |
| PRINTS | PR00688 | Xylose isomerase signature | IPR001998 | Xylose isomerase | 155 | 174 | 9.6E-86 |
| Hamap | MF_00455 | Xylose isomerase [xylA]. | IPR001998 | Xylose isomerase | 10 | 436 | 51.673 |
| PRINTS | PR00688 | Xylose isomerase signature | IPR001998 | Xylose isomerase | 132 | 152 | 9.6E-86 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.