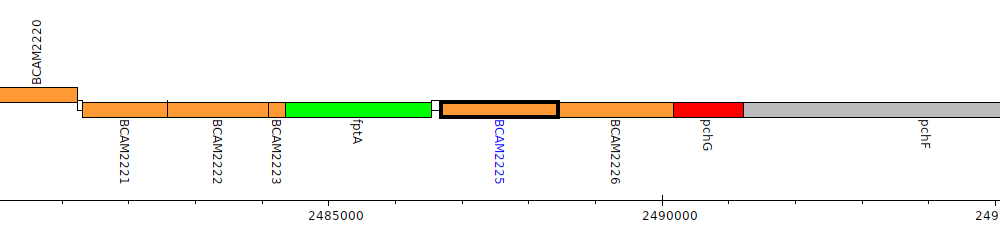
Burkholderia cenocepacia J2315, BCAM2225
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0033212 | iron import into cell | Inferred from Genomic Context | ECO:0000177 genomic context evidence |
||
| Molecular Function | GO:0042626 | ATPase-coupled transmembrane transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS50929
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0042626 | ATPase-coupled transmembrane transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS50929
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016021 | integral component of membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PS50929
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016887 | ATPase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS50893
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS50893
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055085 | transmembrane transport |
Inferred from Sequence Model
Term mapped from: InterPro:PS50929
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016021 | integral component of membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PS50929
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016887 | ATPase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS50893
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS50893
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055085 | transmembrane transport |
Inferred from Sequence Model
Term mapped from: InterPro:PS50929
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:1.20.1560.10 | IPR036640 | ABC transporter type 1, transmembrane domain superfamily | 7 | 315 | 6.7E-13 | |
| SUPERFAMILY | SSF90123 | IPR036640 | ABC transporter type 1, transmembrane domain superfamily | 7 | 320 | 1.2E-26 | |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 566 | 588 | - | ||
| ProSiteProfiles | PS50893 | ATP-binding cassette, ABC transporter-type domain profile. | IPR003439 | ABC transporter-like | 331 | 565 | 21.042 |
| SUPERFAMILY | SSF52540 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 330 | 563 | 4.26E-66 | |
| ProSitePatterns | PS00211 | ABC transporters family signature. | IPR017871 | ABC transporter, conserved site | 468 | 482 | - |
| CDD | cd18561 | ABC_6TM_AarD_CydDC_like | 22 | 291 | 6.66497E-7 | ||
| Gene3D | G3DSA:3.40.50.300 | 326 | 568 | 1.1E-69 | |||
| ProSiteProfiles | PS50929 | ABC transporter integral membrane type-1 fused domain profile. | IPR011527 | ABC transporter type 1, transmembrane domain | 23 | 294 | 20.843 |
| SMART | SM00382 | IPR003593 | AAA+ ATPase domain | 356 | 542 | 1.9E-14 | |
| Pfam | PF00005 | ABC transporter | IPR003439 | ABC transporter-like | 347 | 496 | 3.3E-26 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.