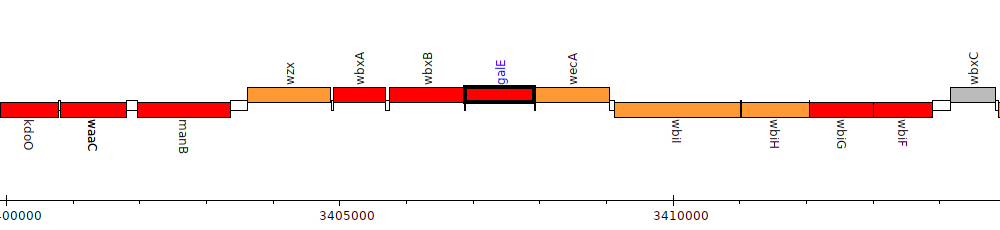
Burkholderia cenocepacia J2315, BCAL3117 (galE)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006012 | galactose metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:cd05247
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003978 | UDP-glucose 4-epimerase activity |
Inferred from Sequence Model
Term mapped from: InterPro:cd05247
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | D-galactose degradation I (Leloir pathway) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | superpathway of UDP-glucose-derived O-antigen building blocks biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj00052 | Galactose metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00052 | Galactose metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj00520 | Amino sugar and nucleotide sugar metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | stachyose degradation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | mycolyl-arabinogalactan-peptidoglycan complex biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | D-galactose detoxification | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | UDP-α-D-galactose biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00520 | Amino sugar and nucleotide sugar metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF51735 | IPR036291 | NAD(P)-binding domain superfamily | 1 | 336 | 2.29E-94 | |
| TIGRFAM | TIGR01179 | galE: UDP-glucose 4-epimerase GalE | IPR005886 | UDP-glucose 4-epimerase | 7 | 337 | 2.5E-132 |
| CDD | cd05247 | UDP_G4E_1_SDR_e | IPR005886 | UDP-glucose 4-epimerase | 6 | 333 | 0.0 |
| Gene3D | G3DSA:3.90.25.10 | 185 | 331 | 3.8E-148 | |||
| Gene3D | G3DSA:3.40.50.720 | 7 | 265 | 3.8E-148 | |||
| Pfam | PF16363 | GDP-mannose 4,6 dehydratase | IPR016040 | NAD(P)-binding domain | 8 | 327 | 6.7E-58 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.