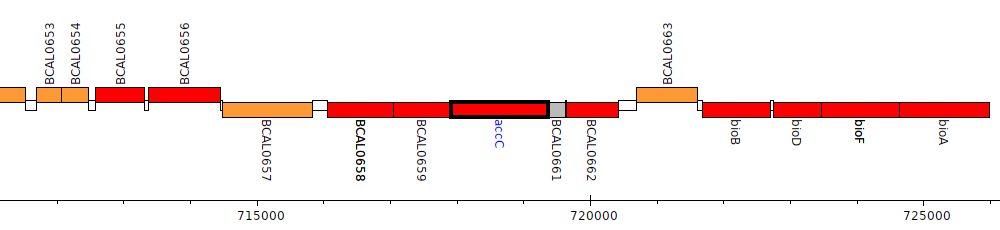
Burkholderia cenocepacia J2315, BCAL0660 (accC)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0046872 | metal ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS50975
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF02786
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj01200 | Carbon metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00640 | Propanoate metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01212 | Fatty acid metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01120 | Microbial metabolism in diverse environments | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00061 | Fatty acid biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00620 | Pyruvate metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| ProSitePatterns | PS00867 | Carbamoyl-phosphate synthase subdomain signature 2. | IPR005479 | Carbamoyl-phosphate synthetase large subunit-like, ATP-binding domain | 300 | 307 | - |
| SMART | SM00878 | IPR005482 | Biotin carboxylase, C-terminal | 350 | 456 | 3.3E-62 | |
| Gene3D | G3DSA:3.30.470.130 | 7 | 460 | 2.6E-192 | |||
| SUPERFAMILY | SSF52440 | IPR016185 | Pre-ATP-grasp domain superfamily | 13 | 127 | 8.63E-45 | |
| ProSiteProfiles | PS50975 | ATP-grasp fold profile. | IPR011761 | ATP-grasp fold | 134 | 331 | 44.551 |
| SUPERFAMILY | SSF56059 | 97 | 366 | 2.65E-68 | |||
| Pfam | PF02786 | Carbamoyl-phosphate synthase L chain, ATP binding domain | IPR005479 | Carbamoyl-phosphate synthetase large subunit-like, ATP-binding domain | 129 | 336 | 4.6E-76 |
| Pfam | PF02785 | Biotin carboxylase C-terminal domain | IPR005482 | Biotin carboxylase, C-terminal | 350 | 456 | 1.3E-37 |
| SUPERFAMILY | SSF51246 | IPR011054 | Rudiment single hybrid motif | 345 | 460 | 3.24E-41 | |
| ProSitePatterns | PS00866 | Carbamoyl-phosphate synthase subdomain signature 1. | IPR005479 | Carbamoyl-phosphate synthetase large subunit-like, ATP-binding domain | 167 | 181 | - |
| ProSiteProfiles | PS50979 | Biotin carboxylation domain profile. | IPR011764 | Biotin carboxylation domain | 15 | 460 | 53.288 |
| Pfam | PF00289 | Biotin carboxylase, N-terminal domain | IPR005481 | Biotin carboxylase-like, N-terminal domain | 16 | 124 | 1.1E-42 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.