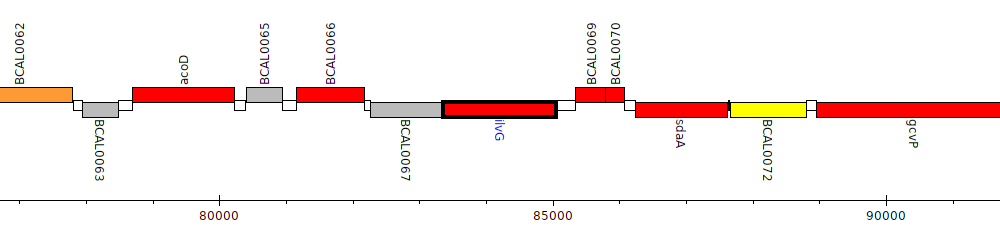
Burkholderia cenocepacia J2315, BCAL0068 (ilvG)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF02775
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0030976 | thiamine pyrophosphate binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS00187
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000287 | magnesium ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS00187
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00650 | Butanoate metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01230 | Biosynthesis of amino acids | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00290 | Valine, leucine and isoleucine biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01210 | 2-Oxocarboxylic acid metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00770 | Pantothenate and CoA biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00660 | C5-Branched dibasic acid metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF02776 | Thiamine pyrophosphate enzyme, N-terminal TPP binding domain | IPR012001 | Thiamine pyrophosphate enzyme, N-terminal TPP-binding domain | 13 | 181 | 3.1E-52 |
| ProSitePatterns | PS00187 | Thiamine pyrophosphate enzymes signature. | IPR000399 | TPP-binding enzyme, conserved site | 431 | 450 | - |
| SUPERFAMILY | SSF52518 | IPR029061 | Thiamin diphosphate-binding fold | 376 | 550 | 1.23E-56 | |
| SUPERFAMILY | SSF52467 | IPR029035 | DHS-like NAD/FAD-binding domain superfamily | 165 | 365 | 6.08E-38 | |
| SUPERFAMILY | SSF52518 | IPR029061 | Thiamin diphosphate-binding fold | 7 | 186 | 7.25E-52 | |
| Gene3D | G3DSA:3.40.50.1220 | 193 | 364 | 6.3E-35 | |||
| CDD | cd07035 | TPP_PYR_POX_like | 17 | 172 | 1.86239E-46 | ||
| Pfam | PF02775 | Thiamine pyrophosphate enzyme, C-terminal TPP binding domain | IPR011766 | Thiamine pyrophosphate enzyme, C-terminal TPP-binding | 395 | 541 | 1.7E-42 |
| Gene3D | G3DSA:3.40.50.970 | 4 | 192 | 2.7E-60 | |||
| CDD | cd00568 | TPP_enzymes | 377 | 543 | 9.14663E-68 | ||
| Pfam | PF00205 | Thiamine pyrophosphate enzyme, central domain | IPR012000 | Thiamine pyrophosphate enzyme, central domain | 201 | 333 | 9.0E-28 |
| Gene3D | G3DSA:3.40.50.970 | 370 | 565 | 1.7E-55 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.