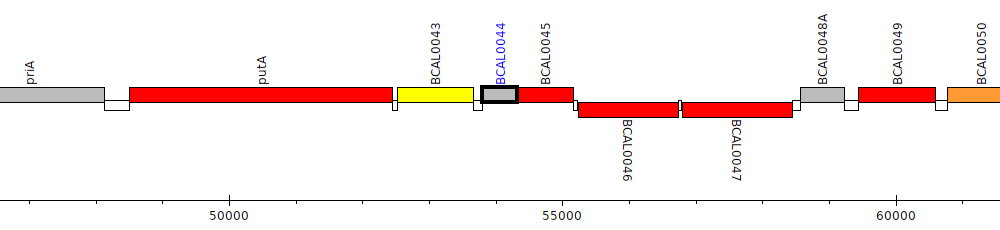
Burkholderia cenocepacia J2315, BCAL0044
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0043565 | sequence-specific DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF48295
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF01527
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004803 | transposase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF01527
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006313 | transposition, DNA-mediated |
Inferred from Sequence Model
Term mapped from: InterPro:PF01527
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF48295 | IPR010921 | Trp repressor/replication initiator | 43 | 116 | 3.77E-17 | |
| SUPERFAMILY | SSF48295 | IPR010921 | Trp repressor/replication initiator | 3 | 54 | 1.31E-11 | |
| Pfam | PF01527 | Transposase | IPR002514 | Transposase IS3/IS911family | 2 | 46 | 4.5E-8 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 108 | 137 | - | ||
| Pfam | PF13518 | Helix-turn-helix domain | 62 | 115 | 2.2E-12 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.