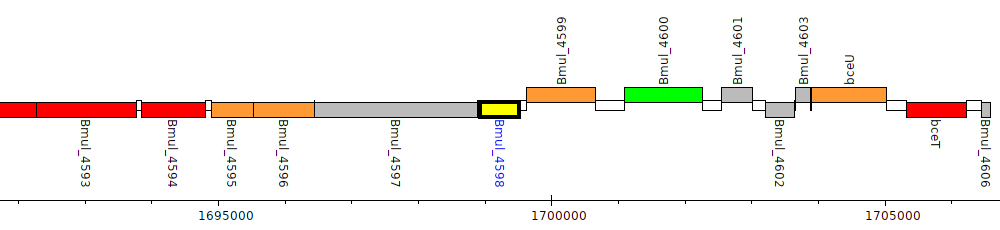
Burkholderia multivorans ATCC 17616, Bmul_4598
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF04879
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0051539 | 4 iron, 4 sulfur cluster binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS00551
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:PF04879
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bmu00680 | Methane metabolism - Burkholderia multivorans ATCC 17616 (JGI) | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bmu01200 | Carbon metabolism - Burkholderia multivorans ATCC 17616 (JGI) | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bmu01120 | Microbial metabolism in diverse environments - Burkholderia multivorans ATCC 17616 (JGI) | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bmu01100 | Metabolic pathways - Burkholderia multivorans ATCC 17616 (JGI) | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bmu00630 | Glyoxylate and dicarboxylate metabolism - Burkholderia multivorans ATCC 17616 (JGI) | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00384 | Molybdopterin oxidoreductase | IPR006656 | Molybdopterin oxidoreductase | 108 | 184 | 8.7E-6 |
| Gene3D | G3DSA:2.20.25.90 | 45 | 105 | 1.3E-17 | |||
| SMART | SM00926 | IPR006963 | Molybdopterin oxidoreductase, 4Fe-4S domain | 44 | 105 | 8.2E-23 | |
| ProSitePatterns | PS00551 | Prokaryotic molybdopterin oxidoreductases signature 1. | IPR027467 | Molybdopterin oxidoreductase, molybdopterin cofactor binding site | 49 | 67 | - |
| ProSiteProfiles | PS51318 | Twin arginine translocation (Tat) signal profile. | IPR006311 | Twin-arginine translocation pathway, signal sequence | 1 | 34 | 9.465 |
| SUPERFAMILY | SSF53706 | 37 | 196 | 5.37E-54 | |||
| Gene3D | G3DSA:3.40.50.740 | 108 | 196 | 2.1E-33 | |||
| ProSiteProfiles | PS51669 | Prokaryotic molybdopterin oxidoreductases 4Fe-4S domain profile. | IPR006963 | Molybdopterin oxidoreductase, 4Fe-4S domain | 44 | 107 | 25.018 |
| Pfam | PF04879 | Molybdopterin oxidoreductase Fe4S4 domain | IPR006963 | Molybdopterin oxidoreductase, 4Fe-4S domain | 46 | 104 | 1.9E-19 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.