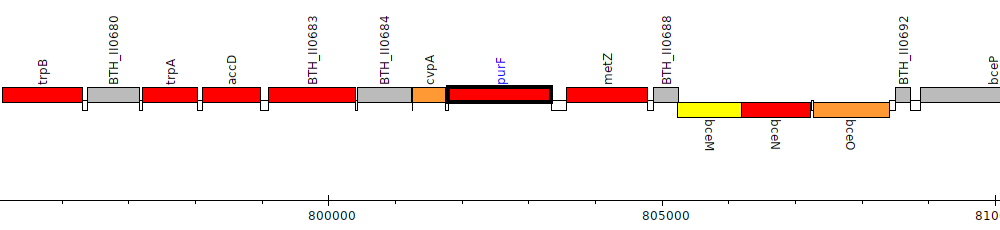
Burkholderia thailandensis E264 ATCC 700388, BTH_II0686 (purF)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0009113 | purine nucleobase biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01134
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009116 | nucleoside metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:cd06223
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009116 | nucleoside metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:cd06223
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004044 | amidophosphoribosyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01134
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009113 | purine nucleobase biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01134
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004044 | amidophosphoribosyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01134
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG (InterPro) | 00250 | Alanine, aspartate and glutamate metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | 5-aminoimidazole ribonucleotide biosynthesis II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bte01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bte01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00230 | Purine metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | 4-amino-2-methyl-5-diphosphomethylpyrimidine biosynthesis (yeast) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bte00230 | Purine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bte01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bte00250 | Alanine, aspartate and glutamate metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | 5-aminoimidazole ribonucleotide biosynthesis I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | superpathway of 5-aminoimidazole ribonucleotide biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.40.50.2020 | 279 | 443 | 4.4E-148 | |||
| CDD | cd06223 | PRTases_typeI | IPR000836 | Phosphoribosyltransferase domain | 279 | 398 | 3.97386E-19 |
| SUPERFAMILY | SSF53271 | IPR029057 | Phosphoribosyltransferase-like | 252 | 482 | 1.28E-70 | |
| TIGRFAM | TIGR01134 | purF: amidophosphoribosyltransferase | IPR005854 | Amidophosphoribosyltransferase | 2 | 466 | 2.9E-153 |
| Hamap | MF_01931 | Amidophosphoribosyltransferase [purF]. | IPR005854 | Amidophosphoribosyltransferase | 1 | 466 | 44.746 |
| Pfam | PF13522 | Glutamine amidotransferase domain | 64 | 210 | 1.3E-14 | ||
| Gene3D | G3DSA:3.60.20.10 | IPR029055 | Nucleophile aminohydrolases, N-terminal | 2 | 414 | 4.4E-148 | |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 486 | 511 | - | ||
| Pfam | PF00156 | Phosphoribosyl transferase domain | IPR000836 | Phosphoribosyltransferase domain | 283 | 395 | 6.3E-11 |
| ProSiteProfiles | PS51278 | Glutamine amidotransferase type 2 domain profile. | IPR017932 | Glutamine amidotransferase type 2 domain | 2 | 236 | 40.743 |
| PIRSF | PIRSF000485 | IPR005854 | Amidophosphoribosyltransferase | 1 | 483 | 9.6E-158 | |
| SUPERFAMILY | SSF56235 | IPR029055 | Nucleophile aminohydrolases, N-terminal | 2 | 259 | 9.44E-69 | |
| CDD | cd00715 | GPATase_N | IPR035584 | Amidophosphoribosyltransferase, N-terminal | 2 | 268 | 2.24006E-134 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.