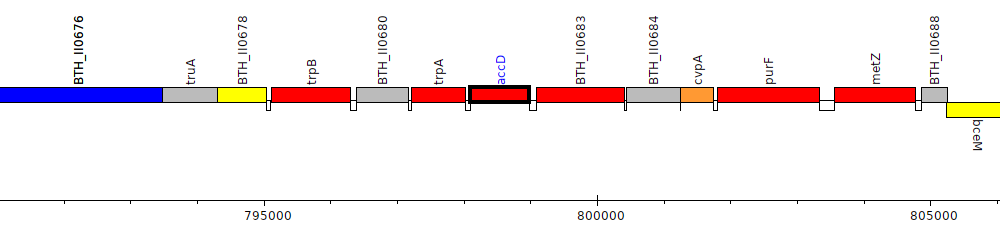
Burkholderia thailandensis E264 ATCC 700388, BTH_II0682 (accD)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Cellular Component | GO:0009317 | acetyl-CoA carboxylase complex |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00515
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006633 | fatty acid biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00515
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0009317 | acetyl-CoA carboxylase complex |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00515
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003989 | acetyl-CoA carboxylase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00515
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003989 | acetyl-CoA carboxylase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00515
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006633 | fatty acid biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00515
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bte01212 | Fatty acid metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bte01200 | Carbon metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bte00640 | Propanoate metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | 3-hydroxypropanoate/4-hydroxybutanate cycle | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | fatty acid biosynthesis initiation I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bte01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | glyoxylate assimilation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | 3-hydroxypropanoate cycle | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bte01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bte00620 | Pyruvate metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bte01120 | Microbial metabolism in diverse environments | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | candicidin biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bte00061 | Fatty acid biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bte01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF17848 | Acetyl-coA carboxylase zinc finger domain | IPR041010 | Acetyl-coA carboxylase zinc finger domain | 28 | 53 | 1.2E-12 |
| PRINTS | PR01070 | Acetyl-CoA carboxylase carboxyl transferase beta subunit signature | IPR000438 | Acetyl-CoA carboxylase carboxyl transferase, beta subunit | 253 | 262 | 8.4E-40 |
| Pfam | PF01039 | Carboxyl transferase domain | IPR034733 | Acetyl-CoA carboxylase | 98 | 242 | 1.1E-13 |
| TIGRFAM | TIGR00515 | accD: acetyl-CoA carboxylase, carboxyl transferase, beta subunit | IPR000438 | Acetyl-CoA carboxylase carboxyl transferase, beta subunit | 1 | 285 | 2.8E-128 |
| PRINTS | PR01070 | Acetyl-CoA carboxylase carboxyl transferase beta subunit signature | IPR000438 | Acetyl-CoA carboxylase carboxyl transferase, beta subunit | 231 | 242 | 8.4E-40 |
| PRINTS | PR01070 | Acetyl-CoA carboxylase carboxyl transferase beta subunit signature | IPR000438 | Acetyl-CoA carboxylase carboxyl transferase, beta subunit | 165 | 183 | 8.4E-40 |
| Gene3D | G3DSA:3.90.226.10 | 27 | 288 | 2.4E-85 | |||
| PRINTS | PR01070 | Acetyl-CoA carboxylase carboxyl transferase beta subunit signature | IPR000438 | Acetyl-CoA carboxylase carboxyl transferase, beta subunit | 202 | 219 | 8.4E-40 |
| SUPERFAMILY | SSF52096 | IPR029045 | ClpP/crotonase-like domain superfamily | 27 | 284 | 1.52E-86 | |
| Hamap | MF_01395 | Acetyl-coenzyme A carboxylase carboxyl transferase subunit beta, chloroplastic [accD]. | IPR000438 | Acetyl-CoA carboxylase carboxyl transferase, beta subunit | 1 | 283 | 38.165 |
| ProSiteProfiles | PS50980 | Acetyl-coenzyme A (CoA) carboxyltransferase N-terminal domain profile. | IPR011762 | Acetyl-coenzyme A carboxyltransferase, N-terminal | 27 | 290 | 106.153 |
| PRINTS | PR01070 | Acetyl-CoA carboxylase carboxyl transferase beta subunit signature | IPR000438 | Acetyl-CoA carboxylase carboxyl transferase, beta subunit | 131 | 145 | 8.4E-40 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.