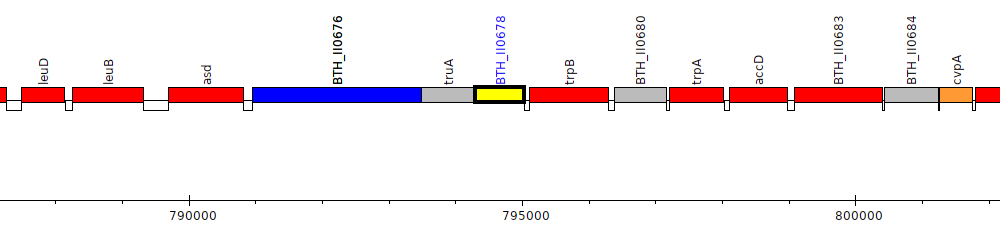
Burkholderia thailandensis E264 ATCC 700388, BTH_II0678
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006568 | tryptophan metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:cd00405
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF51366
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF51366
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004640 | phosphoribosylanthranilate isomerase activity |
Inferred from Sequence Model
Term mapped from: InterPro:cd00405
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004640 | phosphoribosylanthranilate isomerase activity |
Inferred from Sequence Model
Term mapped from: InterPro:cd00405
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006568 | tryptophan metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:cd00405
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bte01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bte00400 | Phenylalanine, tyrosine and tryptophan biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bte01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00400 | Phenylalanine, tyrosine and tryptophan biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bte01230 | Biosynthesis of amino acids | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.20.20.70 | IPR013785 | Aldolase-type TIM barrel | 31 | 245 | 6.0E-64 | |
| Pfam | PF00697 | N-(5'phosphoribosyl)anthranilate (PRA) isomerase | IPR001240 | N-(5'phosphoribosyl) anthranilate isomerase (PRAI) | 36 | 238 | 9.9E-44 |
| SUPERFAMILY | SSF51366 | IPR011060 | Ribulose-phosphate binding barrel | 33 | 241 | 1.57E-54 | |
| CDD | cd00405 | PRAI | IPR001240 | N-(5'phosphoribosyl) anthranilate isomerase (PRAI) | 35 | 240 | 1.14862E-83 |
| Hamap | MF_00135 | N-(5'-phosphoribosyl)anthranilate isomerase [trpF]. | IPR001240 | N-(5'phosphoribosyl) anthranilate isomerase (PRAI) | 33 | 243 | 39.916 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.