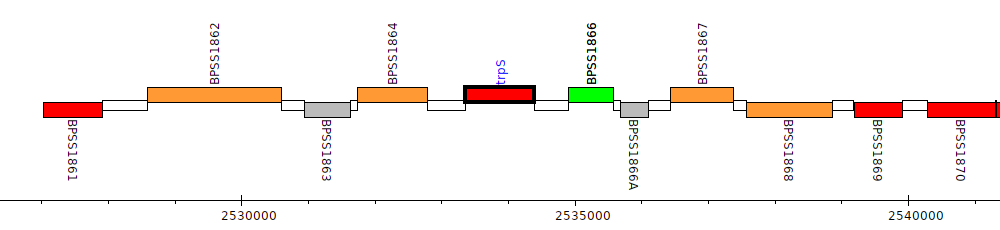
Burkholderia pseudomallei K96243, BPSS1865 (trpS)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0000166 | nucleotide binding |
Inferred from Sequence Model
Term mapped from: InterPro:PR01039
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004812 | aminoacyl-tRNA ligase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00579
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006418 | tRNA aminoacylation for protein translation |
Inferred from Sequence Model
Term mapped from: InterPro:PF00579
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006436 | tryptophanyl-tRNA aminoacylation |
Inferred from Sequence Model
Term mapped from: InterPro:PR01039
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PR01039
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004830 | tryptophan-tRNA ligase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PR01039
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bps00970 | Aminoacyl-tRNA biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00970 | Aminoacyl-tRNA biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF52374 | 13 | 337 | 2.02E-77 | |||
| PRINTS | PR01039 | Tryptophanyl-tRNA synthetase signature | IPR002306 | Tryptophan-tRNA ligase | 75 | 94 | 2.2E-22 |
| PRINTS | PR01039 | Tryptophanyl-tRNA synthetase signature | IPR002306 | Tryptophan-tRNA ligase | 153 | 174 | 2.2E-22 |
| CDD | cd00806 | TrpRS_core | IPR002306 | Tryptophan-tRNA ligase | 15 | 292 | 5.2193E-102 |
| TIGRFAM | TIGR00233 | trpS: tryptophan--tRNA ligase | IPR002306 | Tryptophan-tRNA ligase | 13 | 337 | 4.1E-70 |
| Gene3D | G3DSA:3.40.50.620 | IPR014729 | Rossmann-like alpha/beta/alpha sandwich fold | 15 | 333 | 2.5E-102 | |
| PRINTS | PR01039 | Tryptophanyl-tRNA synthetase signature | IPR002306 | Tryptophan-tRNA ligase | 205 | 215 | 2.2E-22 |
| ProSitePatterns | PS00178 | Aminoacyl-transfer RNA synthetases class-I signature. | IPR001412 | Aminoacyl-tRNA synthetase, class I, conserved site | 20 | 29 | - |
| Pfam | PF00579 | tRNA synthetases class I (W and Y) | IPR002305 | Aminoacyl-tRNA synthetase, class Ic | 10 | 292 | 1.2E-54 |
| Gene3D | G3DSA:1.10.240.10 | 192 | 306 | 2.5E-102 | |||
| PRINTS | PR01039 | Tryptophanyl-tRNA synthetase signature | IPR002306 | Tryptophan-tRNA ligase | 24 | 40 | 2.2E-22 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.