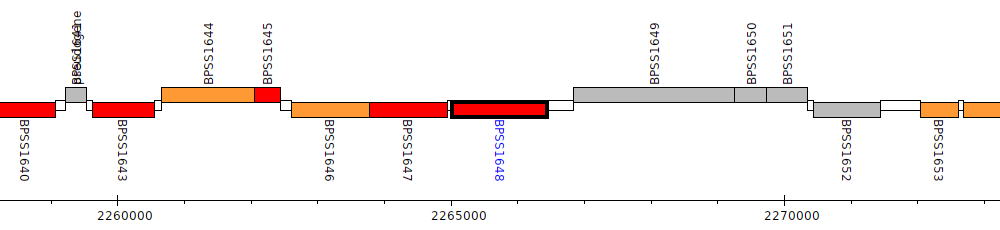
Burkholderia pseudomallei K96243, BPSS1648
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0000160 | phosphorelay signal transduction system |
Inferred from Sequence Model
Term mapped from: InterPro:PS50110
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd00077 | HDc | IPR003607 | HD/PDEase domain | 236 | 365 | 2.1625E-11 |
| Gene3D | G3DSA:3.40.50.2300 | 36 | 176 | 1.1E-35 | |||
| SMART | SM00448 | IPR001789 | Signal transduction response regulator, receiver domain | 39 | 151 | 9.4E-29 | |
| SUPERFAMILY | SSF52172 | IPR011006 | CheY-like superfamily | 41 | 196 | 8.11E-36 | |
| ProSiteProfiles | PS51832 | HD-GYP domain profile. | IPR037522 | HD-GYP domain | 210 | 406 | 30.277 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 1 | 18 | - | ||
| Gene3D | G3DSA:1.10.3210.10 | 207 | 405 | 8.9E-43 | |||
| Pfam | PF00072 | Response regulator receiver domain | IPR001789 | Signal transduction response regulator, receiver domain | 41 | 151 | 1.6E-24 |
| SUPERFAMILY | SSF109604 | 236 | 394 | 1.14E-14 | |||
| Pfam | PF13487 | HD domain | 285 | 346 | 1.2E-18 | ||
| ProSiteProfiles | PS50110 | Response regulatory domain profile. | IPR001789 | Signal transduction response regulator, receiver domain | 40 | 155 | 35.113 |
| CDD | cd00156 | REC | IPR001789 | Signal transduction response regulator, receiver domain | 42 | 155 | 1.49816E-29 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 1 | 32 | - |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.