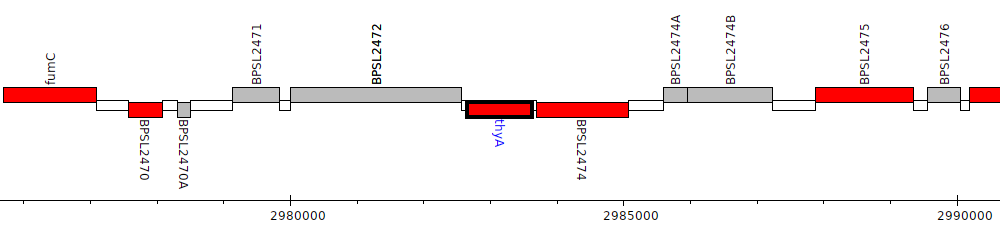
Burkholderia pseudomallei K96243, BPSL2473 (thyA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006231 | dTMP biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00008
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004799 | thymidylate synthase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00008
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | pyrimidine deoxyribonucleotides <i>de novo</i> biosynthesis I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | pyrimidine deoxyribonucleotides biosynthesis from CTP | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | pyrimidine deoxyribonucleosides salvage | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00240 | Pyrimidine metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | pyrimidine deoxyribonucleotides <i>de novo</i> biosynthesis IV | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | folate transformations II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00670 | One carbon pool by folate | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bps00670 | One carbon pool by folate | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | pyrimidine deoxyribonucleotides <i>de novo</i> biosynthesis II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bps00240 | Pyrimidine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00108 | Thymidylate synthase family signature | IPR000398 | Thymidylate synthase | 161 | 180 | 6.8E-26 |
| SUPERFAMILY | SSF55831 | IPR036926 | Thymidylate synthase/dCMP hydroxymethylase superfamily | 1 | 323 | 1.13E-94 | |
| TIGRFAM | TIGR03284 | thym_sym: thymidylate synthase | IPR000398 | Thymidylate synthase | 153 | 323 | 2.8E-34 |
| Hamap | MF_00008 | Thymidylate synthase [thyA]. | IPR000398 | Thymidylate synthase | 1 | 323 | 33.705 |
| Gene3D | G3DSA:3.30.572.10 | IPR036926 | Thymidylate synthase/dCMP hydroxymethylase superfamily | 1 | 323 | 2.0E-124 | |
| PRINTS | PR00108 | Thymidylate synthase family signature | IPR000398 | Thymidylate synthase | 187 | 202 | 6.8E-26 |
| PRINTS | PR00108 | Thymidylate synthase family signature | IPR000398 | Thymidylate synthase | 42 | 63 | 6.8E-26 |
| CDD | cd00351 | TS_Pyrimidine_HMase | IPR023451 | Thymidylate synthase/dCMP hydroxymethylase domain | 3 | 265 | 1.02991E-82 |
| PRINTS | PR00108 | Thymidylate synthase family signature | IPR000398 | Thymidylate synthase | 245 | 262 | 6.8E-26 |
| Pfam | PF00303 | Thymidylate synthase | IPR023451 | Thymidylate synthase/dCMP hydroxymethylase domain | 2 | 323 | 5.5E-83 |
| PRINTS | PR00108 | Thymidylate synthase family signature | IPR000398 | Thymidylate synthase | 207 | 233 | 6.8E-26 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.