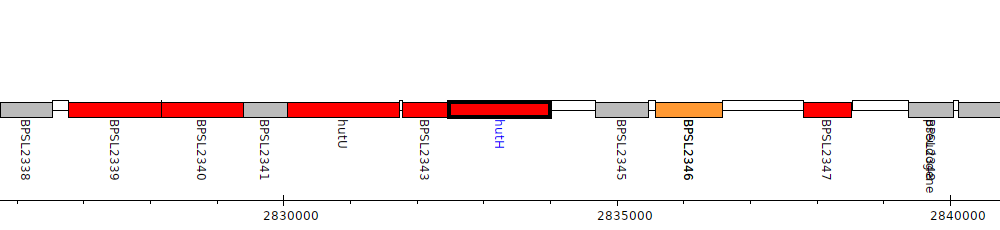
Burkholderia pseudomallei K96243, BPSL2344 (hutH)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Cellular Component | GO:0005737 | cytoplasm |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00229
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016841 | ammonia-lyase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS00488
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004397 | histidine ammonia-lyase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00229
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF48557
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006548 | histidine catabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00229
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bps00340 | Histidine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00340 | Histidine metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | L-histidine degradation III | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bps01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | L-histidine degradation II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:1.10.275.10 | IPR024083 | Fumarase/histidase, N-terminal | 1 | 195 | 4.1E-63 | |
| ProSitePatterns | PS00488 | Phenylalanine and histidine ammonia-lyases signature. | IPR022313 | Phenylalanine/histidine ammonia-lyases, active site | 137 | 153 | - |
| TIGRFAM | TIGR01225 | hutH: histidine ammonia-lyase | IPR005921 | Histidine ammonia-lyase | 2 | 498 | 2.3E-209 |
| Gene3D | G3DSA:1.20.200.10 | 196 | 502 | 1.3E-119 | |||
| CDD | cd00332 | PAL-HAL | IPR001106 | Aromatic amino acid lyase | 9 | 450 | 0.0 |
| Hamap | MF_00229 | Histidine ammonia-lyase [hutH]. | IPR005921 | Histidine ammonia-lyase | 1 | 501 | 44.229 |
| SUPERFAMILY | SSF48557 | IPR008948 | L-Aspartase-like | 1 | 496 | 9.42E-190 | |
| Pfam | PF00221 | Aromatic amino acid lyase | IPR001106 | Aromatic amino acid lyase | 9 | 468 | 2.7E-186 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.