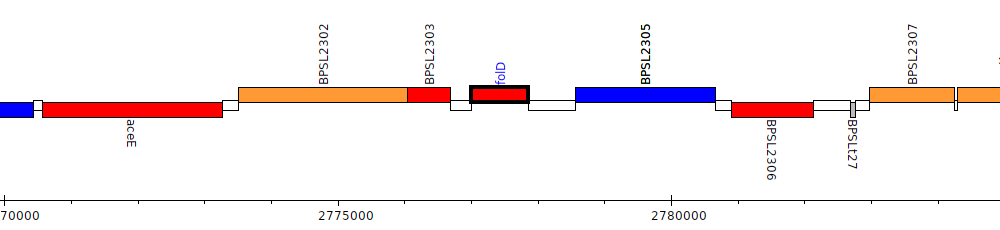
Burkholderia pseudomallei K96243, BPSL2304 (folD)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:PR00085
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS00767
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004488 | methylenetetrahydrofolate dehydrogenase (NADP+) activity |
Inferred from Sequence Model
Term mapped from: InterPro:PR00085
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS00767
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:PR00085
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004488 | methylenetetrahydrofolate dehydrogenase (NADP+) activity |
Inferred from Sequence Model
Term mapped from: InterPro:PR00085
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG (InterPro) | 00720 | Carbon fixation pathways in prokaryotes | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG (InterPro) | 00670 | One carbon pool by folate | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bps01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps01200 | Carbon metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | purine nucleobases degradation II (anaerobic) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | folate transformations II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | formaldehyde oxidation VII (THF pathway) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bps00670 | One carbon pool by folate | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | folate transformations I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | L-histidine degradation III | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | tetrahydrofolate salvage from 5,10-methenyltetrahydrofolate | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | formate assimilation into 5,10-methylenetetrahydrofolate | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bps01120 | Microbial metabolism in diverse environments | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF02882 | Tetrahydrofolate dehydrogenase/cyclohydrolase, NAD(P)-binding domain | IPR020631 | Tetrahydrofolate dehydrogenase/cyclohydrolase, NAD(P)-binding domain | 123 | 281 | 1.3E-70 |
| PRINTS | PR00085 | Tetrahydrofolate dehydrogenase/cyclohydrolase family signature | IPR000672 | Tetrahydrofolate dehydrogenase/cyclohydrolase | 203 | 232 | 2.4E-85 |
| PRINTS | PR00085 | Tetrahydrofolate dehydrogenase/cyclohydrolase family signature | IPR000672 | Tetrahydrofolate dehydrogenase/cyclohydrolase | 74 | 101 | 2.4E-85 |
| ProSitePatterns | PS00766 | Tetrahydrofolate dehydrogenase/cyclohydrolase signature 1. | IPR020867 | Tetrahydrofolate dehydrogenase/cyclohydrolase, conserved site | 75 | 100 | - |
| PRINTS | PR00085 | Tetrahydrofolate dehydrogenase/cyclohydrolase family signature | IPR000672 | Tetrahydrofolate dehydrogenase/cyclohydrolase | 33 | 55 | 2.4E-85 |
| SUPERFAMILY | SSF53223 | 1 | 121 | 6.43E-40 | |||
| ProSitePatterns | PS00767 | Tetrahydrofolate dehydrogenase/cyclohydrolase signature 2. | IPR020867 | Tetrahydrofolate dehydrogenase/cyclohydrolase, conserved site | 259 | 267 | - |
| PRINTS | PR00085 | Tetrahydrofolate dehydrogenase/cyclohydrolase family signature | IPR000672 | Tetrahydrofolate dehydrogenase/cyclohydrolase | 238 | 254 | 2.4E-85 |
| SUPERFAMILY | SSF51735 | IPR036291 | NAD(P)-binding domain superfamily | 122 | 280 | 4.04E-53 | |
| Gene3D | G3DSA:3.40.50.720 | 6 | 258 | 3.9E-112 | |||
| PRINTS | PR00085 | Tetrahydrofolate dehydrogenase/cyclohydrolase family signature | IPR000672 | Tetrahydrofolate dehydrogenase/cyclohydrolase | 154 | 174 | 2.4E-85 |
| Pfam | PF00763 | Tetrahydrofolate dehydrogenase/cyclohydrolase, catalytic domain | IPR020630 | Tetrahydrofolate dehydrogenase/cyclohydrolase, catalytic domain | 5 | 120 | 4.3E-40 |
| PRINTS | PR00085 | Tetrahydrofolate dehydrogenase/cyclohydrolase family signature | IPR000672 | Tetrahydrofolate dehydrogenase/cyclohydrolase | 255 | 273 | 2.4E-85 |
| Hamap | MF_01576 | Bifunctional protein FolD [folD]. | IPR000672 | Tetrahydrofolate dehydrogenase/cyclohydrolase | 3 | 282 | 38.967 |
| PRINTS | PR00085 | Tetrahydrofolate dehydrogenase/cyclohydrolase family signature | IPR000672 | Tetrahydrofolate dehydrogenase/cyclohydrolase | 109 | 130 | 2.4E-85 |
| CDD | cd01080 | NAD_bind_m-THF_DH_Cyclohyd | IPR020631 | Tetrahydrofolate dehydrogenase/cyclohydrolase, NAD(P)-binding domain | 115 | 275 | 1.27841E-90 |
| Gene3D | G3DSA:3.40.50.10860 | 8 | 276 | 3.9E-112 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.