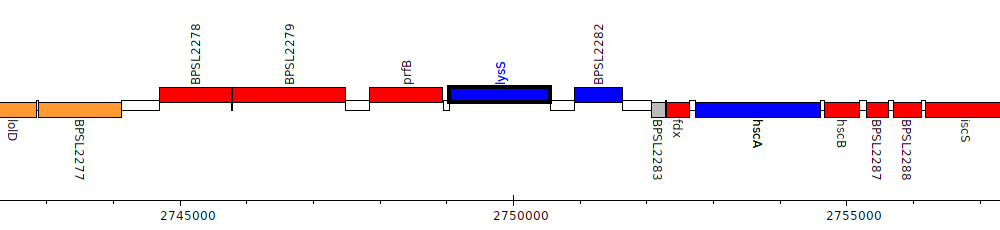
Burkholderia pseudomallei K96243, BPSL2281 (lysS)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006418 | tRNA aminoacylation for protein translation |
Inferred from Sequence Model
Term mapped from: InterPro:PF00152
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:cd00775
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000166 | nucleotide binding |
Inferred from Sequence Model
Term mapped from: InterPro:cd00775
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004824 | lysine-tRNA ligase activity |
Inferred from Sequence Model
Term mapped from: InterPro:cd00775
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004812 | aminoacyl-tRNA ligase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00152
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000166 | nucleotide binding |
Inferred from Sequence Model
Term mapped from: InterPro:cd00775
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006418 | tRNA aminoacylation for protein translation |
Inferred from Sequence Model
Term mapped from: InterPro:PF00152
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0005737 | cytoplasm |
Inferred from Sequence Model
Term mapped from: InterPro:cd00775
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004824 | lysine-tRNA ligase activity |
Inferred from Sequence Model
Term mapped from: InterPro:cd00775
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003676 | nucleic acid binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF01336
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0005737 | cytoplasm |
Inferred from Sequence Model
Term mapped from: InterPro:cd00775
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006430 | lysyl-tRNA aminoacylation |
Inferred from Sequence Model
Term mapped from: InterPro:cd00775
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003676 | nucleic acid binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF01336
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004812 | aminoacyl-tRNA ligase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00152
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006430 | lysyl-tRNA aminoacylation |
Inferred from Sequence Model
Term mapped from: InterPro:cd00775
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:cd00775
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG (InterPro) | 00970 | Aminoacyl-tRNA biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bps00970 | Aminoacyl-tRNA biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00982 | Lysyl-tRNA synthetase signature | IPR018149 | Lysyl-tRNA synthetase, class II, C-terminal | 214 | 230 | 2.2E-38 |
| ProSiteProfiles | PS50862 | Aminoacyl-transfer RNA synthetases class-II family profile. | IPR006195 | Aminoacyl-tRNA synthetase, class II | 183 | 502 | 28.154 |
| TIGRFAM | TIGR00499 | lysS_bact: lysine--tRNA ligase | IPR002313 | Lysine-tRNA ligase, class II | 15 | 506 | 7.7E-198 |
| Gene3D | G3DSA:2.40.50.140 | 15 | 153 | 1.0E-40 | |||
| Pfam | PF00152 | tRNA synthetases class II (D, K and N) | IPR004364 | Aminoacyl-tRNA synthetase, class II (D/K/N) | 161 | 504 | 9.5E-83 |
| Pfam | PF01336 | OB-fold nucleic acid binding domain | IPR004365 | OB-fold nucleic acid binding domain, AA-tRNA synthetase-type | 68 | 145 | 4.1E-14 |
| PRINTS | PR00982 | Lysyl-tRNA synthetase signature | IPR018149 | Lysyl-tRNA synthetase, class II, C-terminal | 198 | 208 | 2.2E-38 |
| PRINTS | PR00982 | Lysyl-tRNA synthetase signature | IPR018149 | Lysyl-tRNA synthetase, class II, C-terminal | 243 | 256 | 2.2E-38 |
| PRINTS | PR00982 | Lysyl-tRNA synthetase signature | IPR018149 | Lysyl-tRNA synthetase, class II, C-terminal | 261 | 278 | 2.2E-38 |
| Hamap | MF_00252 | Lysine--tRNA ligase [lysS]. | IPR002313 | Lysine-tRNA ligase, class II | 21 | 507 | 43.93 |
| SUPERFAMILY | SSF50249 | IPR012340 | Nucleic acid-binding, OB-fold | 14 | 154 | 4.29E-41 | |
| CDD | cd00775 | LysRS_core | IPR018149 | Lysyl-tRNA synthetase, class II, C-terminal | 177 | 505 | 0.0 |
| CDD | cd04322 | LysRS_N | 67 | 173 | 1.21421E-56 | ||
| Gene3D | G3DSA:3.30.930.10 | 161 | 505 | 6.6E-133 | |||
| PRINTS | PR00982 | Lysyl-tRNA synthetase signature | IPR018149 | Lysyl-tRNA synthetase, class II, C-terminal | 389 | 405 | 2.2E-38 |
| SUPERFAMILY | SSF55681 | 161 | 505 | 4.42E-116 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.