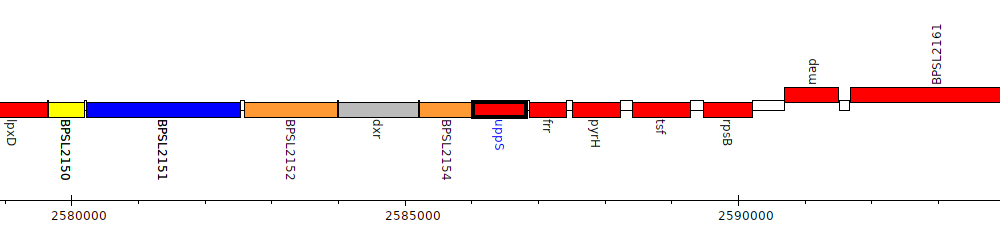
Burkholderia pseudomallei K96243, BPSL2155 (uppS)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016765 | transferase activity, transferring alkyl or aryl (other than methyl) groups |
Inferred from Sequence Model
Term mapped from: InterPro:SSF64005
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bps01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps00900 | Terpenoid backbone biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| TIGRFAM | TIGR00055 | uppS: di-trans,poly-cis-decaprenylcistransferase | IPR001441 | Decaprenyl diphosphate synthase-like | 16 | 242 | 5.5E-89 |
| CDD | cd00475 | Cis_IPPS | IPR001441 | Decaprenyl diphosphate synthase-like | 19 | 234 | 4.13723E-129 |
| SUPERFAMILY | SSF64005 | IPR036424 | Decaprenyl diphosphate synthase-like superfamily | 12 | 237 | 1.44E-86 | |
| Hamap | MF_01139 | Ditrans,polycis-undecaprenyl-diphosphate synthase ((2E,6E)-farnesyl-diphosphate specific) [uppS]. | IPR001441 | Decaprenyl diphosphate synthase-like | 16 | 243 | 40.443 |
| ProSitePatterns | PS01066 | Undecaprenyl pyrophosphate synthase family signature. | IPR018520 | Di-trans-poly-cis-decaprenylcistransferase-like, conserved site | 188 | 205 | - |
| Pfam | PF01255 | Putative undecaprenyl diphosphate synthase | IPR001441 | Decaprenyl diphosphate synthase-like | 23 | 242 | 2.8E-80 |
| Gene3D | G3DSA:3.40.1180.10 | IPR036424 | Decaprenyl diphosphate synthase-like superfamily | 3 | 243 | 6.5E-93 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.