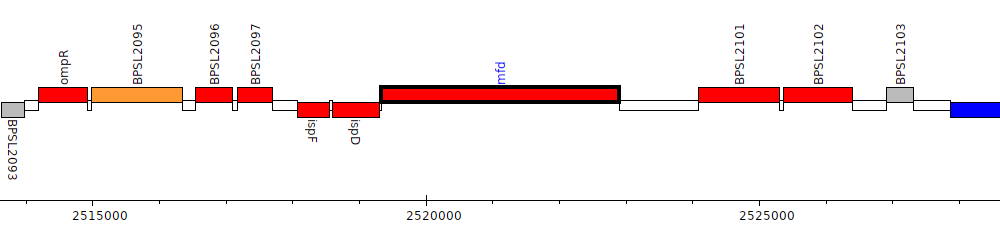
Burkholderia pseudomallei K96243, BPSL2100 (mfd)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003684 | damaged DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00580
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003684 | damaged DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00580
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00270
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003676 | nucleic acid binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00270
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006281 | DNA repair |
Inferred from Sequence Model
Term mapped from: InterPro:SM00982
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003676 | nucleic acid binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00270
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00270
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006281 | DNA repair |
Inferred from Sequence Model
Term mapped from: InterPro:SM00982
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bps03420 | Nucleotide excision repair | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00270 | DEAD/DEAH box helicase | IPR011545 | DEAD/DEAH box helicase domain | 637 | 794 | 7.4E-17 |
| SMART | SM01058 | IPR003711 | CarD-like/TRCF domain | 511 | 608 | 7.3E-50 | |
| Gene3D | G3DSA:3.40.50.300 | 574 | 813 | 9.7E-232 | |||
| Gene3D | G3DSA:3.40.50.11140 | 387 | 484 | 1.2E-14 | |||
| SUPERFAMILY | SSF52540 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 47 | 383 | 8.85E-94 | |
| CDD | cd18810 | SF2_C_TRCF | 820 | 970 | 3.46483E-76 | ||
| Gene3D | G3DSA:3.40.50.300 | 510 | 1022 | 9.7E-232 | |||
| CDD | cd17991 | DEXHc_TRCF | 621 | 813 | 5.88096E-117 | ||
| Gene3D | G3DSA:2.30.30.840 | IPR036101 | CarD-like/TRCF domain superfamily | 515 | 573 | 9.7E-232 | |
| TIGRFAM | TIGR00580 | mfd: transcription-repair coupling factor | IPR004576 | Transcription-repair coupling factor | 187 | 1113 | 0.0 |
| SMART | SM00490 | IPR001650 | Helicase, C-terminal | 858 | 942 | 4.4E-17 | |
| SUPERFAMILY | SSF143517 | IPR037235 | TRCF-like, C-terminal domain | 1026 | 1177 | 3.14E-39 | |
| Hamap | MF_00969 | Transcription-repair-coupling factor [mfd]. | IPR004576 | Transcription-repair coupling factor | 75 | 1141 | 27.439 |
| ProSiteProfiles | PS51194 | Superfamilies 1 and 2 helicase C-terminal domain profile. | IPR001650 | Helicase, C-terminal | 820 | 986 | 15.986 |
| SMART | SM00487 | IPR014001 | Helicase superfamily 1/2, ATP-binding domain | 632 | 822 | 8.7E-32 | |
| SMART | SM00982 | IPR005118 | Transcription-repair-coupling factor, C-terminal domain | 1040 | 1140 | 1.3E-45 | |
| SUPERFAMILY | SSF52540 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 388 | 496 | 4.25E-23 | |
| Gene3D | G3DSA:3.90.1150.50 | IPR037235 | TRCF-like, C-terminal domain | 1036 | 1180 | 7.6E-39 | |
| Gene3D | G3DSA:3.30.2060.10 | 168 | 253 | 2.4E-113 | |||
| SUPERFAMILY | SSF52540 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 555 | 815 | 5.93E-59 | |
| Pfam | PF03461 | TRCF domain | IPR005118 | Transcription-repair-coupling factor, C-terminal domain | 1041 | 1133 | 2.2E-25 |
| SUPERFAMILY | SSF52540 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 815 | 1022 | 5.45E-48 | |
| ProSiteProfiles | PS51192 | Superfamilies 1 and 2 helicase ATP-binding type-1 domain profile. | IPR014001 | Helicase superfamily 1/2, ATP-binding domain | 650 | 811 | 22.958 |
| Gene3D | G3DSA:3.40.50.11180 | 60 | 367 | 2.4E-113 | |||
| Pfam | PF00271 | Helicase conserved C-terminal domain | IPR001650 | Helicase, C-terminal | 838 | 941 | 6.5E-17 |
| Pfam | PF17757 | UvrB interaction domain | IPR041471 | UvrB, interaction domain | 169 | 257 | 7.3E-18 |
| Pfam | PF02559 | CarD-like/TRCF domain | IPR003711 | CarD-like/TRCF domain | 512 | 607 | 1.1E-25 |
| SUPERFAMILY | SSF141259 | IPR036101 | CarD-like/TRCF domain superfamily | 503 | 580 | 1.96E-23 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.