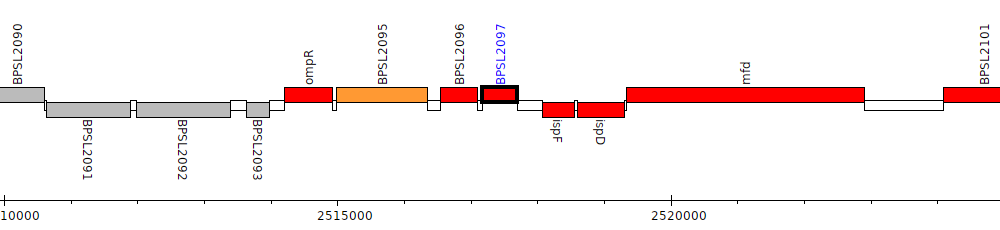
Burkholderia pseudomallei K96243, BPSL2097
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01676
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0051920 | peroxiredoxin activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01676
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0032843 | hydroperoxide reductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01676
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006979 | response to oxidative stress |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01676
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG (InterPro) | 00480 | Glutathione metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF02627 | Carboxymuconolactone decarboxylase family | IPR003779 | Carboxymuconolactone decarboxylase-like | 94 | 169 | 6.9E-13 |
| SUPERFAMILY | SSF69118 | IPR029032 | AhpD-like | 3 | 172 | 2.76E-38 | |
| Gene3D | G3DSA:1.20.1290.10 | IPR029032 | AhpD-like | 3 | 175 | 2.3E-48 | |
| Hamap | MF_01676 | Alkyl hydroperoxide reductase AhpD [ahpD]. | IPR004674 | Alkylhydroperoxidase AhpD | 1 | 175 | 75.364 |
| TIGRFAM | TIGR00778 | ahpD_dom: alkylhydroperoxidase AhpD family core domain | IPR004675 | Alkylhydroperoxidase AhpD core | 111 | 159 | 1.5E-7 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.