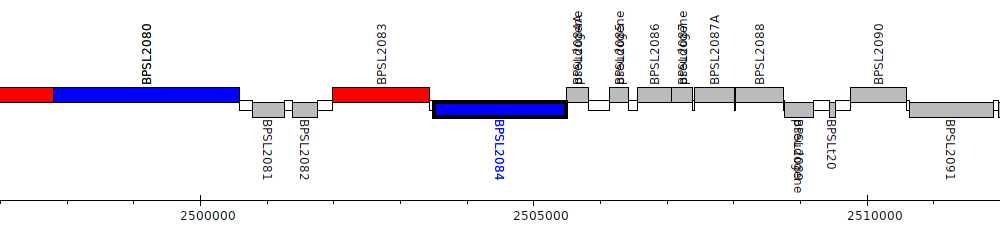
Burkholderia pseudomallei K96243, BPSL2084
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006355 | regulation of transcription, DNA-templated |
Inferred from Sequence Model
Term mapped from: InterPro:PS50937
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006355 | regulation of transcription, DNA-templated |
Inferred from Sequence Model
Term mapped from: InterPro:PS50937
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008168 | methyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF04072
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008168 | methyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF04072
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0032259 | methylation |
Inferred from Sequence Model
Term mapped from: InterPro:PF04072
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS50937
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS50937
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0032259 | methylation |
Inferred from Sequence Model
Term mapped from: InterPro:PF04072
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00040 | MerR bacterial regulatory protein HTH signature | IPR000551 | MerR-type HTH domain | 68 | 88 | 1.3E-7 |
| Pfam | PF04072 | Leucine carboxyl methyltransferase | IPR007213 | Methyltransferase Ppm1/Ppm2/Tcmp | 387 | 563 | 1.5E-45 |
| Coils | Coil | 107 | 127 | - | |||
| Pfam | PF13411 | MerR HTH family regulatory protein | IPR000551 | MerR-type HTH domain | 33 | 99 | 6.8E-15 |
| SUPERFAMILY | SSF53335 | IPR029063 | S-adenosyl-L-methionine-dependent methyltransferase | 385 | 645 | 1.79E-63 | |
| Gene3D | G3DSA:3.40.50.150 | 375 | 644 | 9.9E-59 | |||
| PRINTS | PR00040 | MerR bacterial regulatory protein HTH signature | IPR000551 | MerR-type HTH domain | 44 | 57 | 1.3E-7 |
| ProSitePatterns | PS00552 | MerR-type HTH domain signature. | IPR000551 | MerR-type HTH domain | 35 | 57 | - |
| SMART | SM00422 | IPR000551 | MerR-type HTH domain | 32 | 101 | 7.9E-23 | |
| SUPERFAMILY | SSF46955 | IPR009061 | Putative DNA-binding domain superfamily | 32 | 138 | 1.58E-26 | |
| PRINTS | PR00040 | MerR bacterial regulatory protein HTH signature | IPR000551 | MerR-type HTH domain | 33 | 44 | 1.3E-7 |
| TIGRFAM | TIGR00027 | mthyl_TIGR00027: methyltransferase, TIGR00027 family | IPR011610 | S-adenosyl-L-methionine-dependent methyltransferase, putative | 386 | 638 | 3.1E-56 |
| ProSiteProfiles | PS50937 | MerR-type HTH domain profile. | IPR000551 | MerR-type HTH domain | 31 | 100 | 34.768 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.