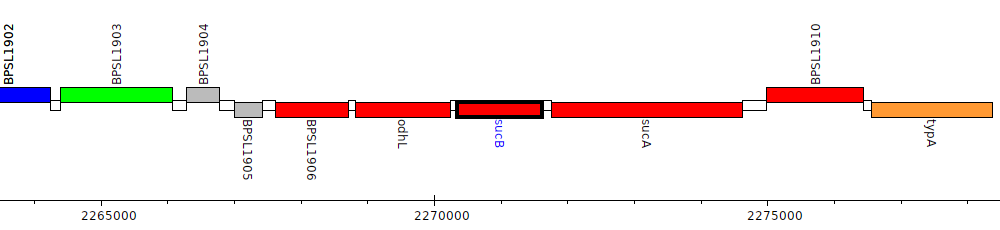
Burkholderia pseudomallei K96243, BPSL1908 (sucB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Cellular Component | GO:0045252 | oxoglutarate dehydrogenase complex |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01347
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006099 | tricarboxylic acid cycle |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01347
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004149 | dihydrolipoyllysine-residue succinyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01347
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016746 | transferase activity, transferring acyl groups |
Inferred from Sequence Model
Term mapped from: InterPro:PF02817
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006099 | tricarboxylic acid cycle |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01347
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004149 | dihydrolipoyllysine-residue succinyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01347
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0045252 | oxoglutarate dehydrogenase complex |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01347
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016746 | transferase activity, transferring acyl groups |
Inferred from Sequence Model
Term mapped from: InterPro:PF02817
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG (InterPro) | 00310 | Lysine degradation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG (InterPro) | 00020 | Citrate cycle (TCA cycle) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bps01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps00020 | Citrate cycle (TCA cycle) | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | 2-oxoglutarate decarboxylation to succinyl-CoA | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bps01200 | Carbon metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps01120 | Microbial metabolism in diverse environments | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps00310 | Lysine degradation | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00380 | Tryptophan metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bps01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:4.10.320.10 | IPR036625 | E3-binding domain superfamily | 112 | 159 | 1.2E-10 | |
| SUPERFAMILY | SSF47005 | IPR036625 | E3-binding domain superfamily | 115 | 157 | 1.57E-9 | |
| Gene3D | G3DSA:3.30.559.10 | IPR023213 | Chloramphenicol acetyltransferase-like domain superfamily | 193 | 425 | 2.8E-96 | |
| Pfam | PF00364 | Biotin-requiring enzyme | IPR000089 | Biotin/lipoyl attachment | 5 | 77 | 7.0E-22 |
| SUPERFAMILY | SSF51230 | IPR011053 | Single hybrid motif | 2 | 95 | 2.36E-26 | |
| CDD | cd06849 | lipoyl_domain | 4 | 77 | 3.1263E-27 | ||
| SUPERFAMILY | SSF52777 | 195 | 424 | 5.78E-90 | |||
| Gene3D | G3DSA:2.40.50.100 | 2 | 104 | 2.3E-23 | |||
| ProSiteProfiles | PS50968 | Biotinyl/lipoyl domain profile. | IPR000089 | Biotin/lipoyl attachment | 3 | 78 | 28.484 |
| ProSiteProfiles | PS51826 | Peripheral subunit-binding (PSBD) domain profile. | IPR004167 | Peripheral subunit-binding domain | 121 | 158 | 17.982 |
| Pfam | PF00198 | 2-oxoacid dehydrogenases acyltransferase (catalytic domain) | IPR001078 | 2-oxoacid dehydrogenase acyltransferase, catalytic domain | 195 | 423 | 2.2E-81 |
| ProSitePatterns | PS00189 | 2-oxo acid dehydrogenases acyltransferase component lipoyl binding site. | IPR003016 | 2-oxo acid dehydrogenase, lipoyl-binding site | 28 | 57 | - |
| TIGRFAM | TIGR01347 | sucB: dihydrolipoyllysine-residue succinyltransferase, E2 component of oxoglutarate dehydrogenase (succinyl-transferring) complex | IPR006255 | Dihydrolipoamide succinyltransferase | 4 | 424 | 1.5E-171 |
| Pfam | PF02817 | e3 binding domain | IPR004167 | Peripheral subunit-binding domain | 122 | 155 | 7.9E-12 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.