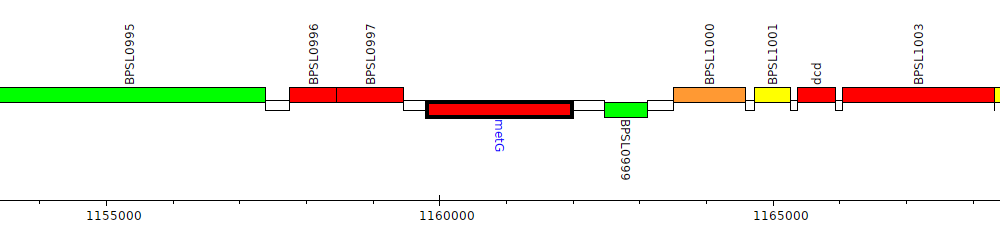
Burkholderia pseudomallei K96243, BPSL0998 (metG)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0000049 | tRNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS50886
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004812 | aminoacyl-tRNA ligase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS00178
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006418 | tRNA aminoacylation for protein translation |
Inferred from Sequence Model
Term mapped from: InterPro:PS00178
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000166 | nucleotide binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00399
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006431 | methionyl-tRNA aminoacylation |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00399
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004825 | methionine-tRNA ligase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00399
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00399
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bps00450 | Selenocompound metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00450 | Selenocompound metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bps00970 | Aminoacyl-tRNA biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00970 | Aminoacyl-tRNA biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| TIGRFAM | TIGR00399 | metG_C_term: methionine--tRNA ligase, beta subunit | IPR004495 | Methionyl-tRNA synthetase, beta subunit, C-terminal | 594 | 725 | 5.2E-40 |
| PRINTS | PR01041 | Methionyl-tRNA synthetase signature | IPR033911 | Methioninyl-tRNA synthetase core domain | 307 | 322 | 1.5E-23 |
| PRINTS | PR01041 | Methionyl-tRNA synthetase signature | IPR033911 | Methioninyl-tRNA synthetase core domain | 268 | 279 | 1.5E-23 |
| PRINTS | PR01041 | Methionyl-tRNA synthetase signature | IPR033911 | Methioninyl-tRNA synthetase core domain | 22 | 35 | 1.5E-23 |
| PRINTS | PR01041 | Methionyl-tRNA synthetase signature | IPR033911 | Methioninyl-tRNA synthetase core domain | 102 | 113 | 1.5E-23 |
| Hamap | MF_00098 | Methionine--tRNA ligase [metG]. | IPR023458 | Methionine-tRNA ligase, type 1 | 18 | 564 | 39.207 |
| SUPERFAMILY | SSF47323 | IPR009080 | Aminoacyl-tRNA synthetase, class Ia, anticodon-binding | 403 | 633 | 1.23E-38 | |
| Gene3D | G3DSA:1.10.730.10 | 406 | 544 | 4.1E-30 | |||
| Pfam | PF09334 | tRNA synthetases class I (M) | IPR015413 | Methionyl/Leucyl tRNA synthetase | 20 | 413 | 3.3E-146 |
| SUPERFAMILY | SSF52374 | 17 | 404 | 5.87E-95 | |||
| TIGRFAM | TIGR00398 | metG: methionine--tRNA ligase | IPR014758 | Methionyl-tRNA synthetase | 20 | 527 | 3.7E-158 |
| Gene3D | G3DSA:2.20.28.20 | IPR029038 | Methionyl-tRNA synthetase, Zn-domain | 154 | 188 | 1.3E-143 | |
| Coils | Coil | 377 | 397 | - | |||
| SUPERFAMILY | SSF50249 | IPR012340 | Nucleic acid-binding, OB-fold | 618 | 724 | 2.69E-30 | |
| Pfam | PF01588 | Putative tRNA binding domain | IPR002547 | tRNA-binding domain | 625 | 722 | 6.1E-23 |
| ProSitePatterns | PS00178 | Aminoacyl-transfer RNA synthetases class-I signature. | IPR001412 | Aminoacyl-tRNA synthetase, class I, conserved site | 27 | 38 | - |
| PRINTS | PR01041 | Methionyl-tRNA synthetase signature | IPR033911 | Methioninyl-tRNA synthetase core domain | 401 | 412 | 1.5E-23 |
| Gene3D | G3DSA:3.40.50.620 | IPR014729 | Rossmann-like alpha/beta/alpha sandwich fold | 20 | 401 | 1.3E-143 | |
| PRINTS | PR01041 | Methionyl-tRNA synthetase signature | IPR033911 | Methioninyl-tRNA synthetase core domain | 54 | 68 | 1.5E-23 |
| Gene3D | G3DSA:2.40.50.140 | 610 | 725 | 4.4E-34 | |||
| CDD | cd00814 | MetRS_core | IPR033911 | Methioninyl-tRNA synthetase core domain | 20 | 381 | 6.32052E-125 |
| CDD | cd07957 | Anticodon_Ia_Met | IPR041872 | Methionyl-tRNA synthetase, anticodon-binding domain | 391 | 523 | 1.71727E-36 |
| CDD | cd02800 | tRNA_bind_EcMetRS_like | IPR004495 | Methionyl-tRNA synthetase, beta subunit, C-terminal | 615 | 725 | 4.38975E-50 |
| ProSiteProfiles | PS50886 | tRNA-binding domain profile. | IPR002547 | tRNA-binding domain | 619 | 725 | 28.483 |
| SUPERFAMILY | SSF57770 | IPR029038 | Methionyl-tRNA synthetase, Zn-domain | 154 | 188 | 4.58E-12 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.