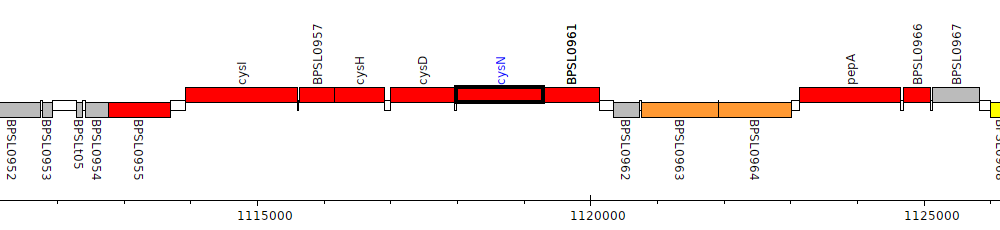
Burkholderia pseudomallei K96243, BPSL0960 (cysN)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003924 | GTPase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PR00315
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006790 | sulfur compound metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02034
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005525 | GTP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PR00315
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG (InterPro) | 00230 | Purine metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bps01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps00230 | Purine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps00920 | Sulfur metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | selenate reduction | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00920 | Sulfur metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG (InterPro) | 00261 | Monobactam biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | assimilatory sulfate reduction III | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bps00450 | Selenocompound metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00450 | Selenocompound metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | sulfite oxidation III | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bps01120 | Microbial metabolism in diverse environments | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | sulfate activation for sulfonation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00315 | GTP-binding elongation factor signature | IPR000795 | Transcription factor, GTP-binding domain | 94 | 104 | 4.2E-13 |
| PRINTS | PR00315 | GTP-binding elongation factor signature | IPR000795 | Transcription factor, GTP-binding domain | 110 | 121 | 4.2E-13 |
| SUPERFAMILY | SSF50447 | IPR009000 | Translation protein, beta-barrel domain superfamily | 233 | 324 | 9.56E-17 | |
| Pfam | PF00009 | Elongation factor Tu GTP binding domain | IPR000795 | Transcription factor, GTP-binding domain | 13 | 205 | 7.8E-38 |
| PRINTS | PR00315 | GTP-binding elongation factor signature | IPR000795 | Transcription factor, GTP-binding domain | 74 | 82 | 4.2E-13 |
| Gene3D | G3DSA:2.40.30.10 | 232 | 326 | 7.2E-21 | |||
| SUPERFAMILY | SSF50465 | IPR009001 | Translation elongation factor EF1A/initiation factor IF2gamma, C-terminal | 331 | 434 | 2.54E-29 | |
| TIGRFAM | TIGR02034 | CysN: sulfate adenylyltransferase, large subunit | IPR011779 | Sulphate adenylyltransferase, large subunit | 13 | 433 | 1.7E-158 |
| CDD | cd04095 | CysN_NoDQ_III | 330 | 433 | 6.59702E-42 | ||
| PRINTS | PR00315 | GTP-binding elongation factor signature | IPR000795 | Transcription factor, GTP-binding domain | 14 | 27 | 4.2E-13 |
| CDD | cd04166 | CysN_ATPS | IPR041757 | Sulfate adenylyltransferase subunit CysN, GTP-binding domain | 14 | 229 | 1.5505E-131 |
| SUPERFAMILY | SSF52540 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 13 | 244 | 1.38E-53 | |
| ProSiteProfiles | PS51722 | Translational (tr)-type guanine nucleotide-binding (G) domain profile. | IPR000795 | Transcription factor, GTP-binding domain | 10 | 232 | 38.12 |
| Gene3D | G3DSA:3.40.50.300 | 4 | 231 | 2.0E-64 | |||
| PRINTS | PR00315 | GTP-binding elongation factor signature | IPR000795 | Transcription factor, GTP-binding domain | 155 | 164 | 4.2E-13 |
| ProSitePatterns | PS00301 | Translational (tr)-type guanine nucleotide-binding (G) domain signature. | IPR031157 | Tr-type G domain, conserved site | 67 | 82 | - |
| CDD | cd03695 | CysN_NodQ_II | 236 | 324 | 1.11377E-33 | ||
| Gene3D | G3DSA:2.40.30.10 | 330 | 434 | 9.0E-33 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.