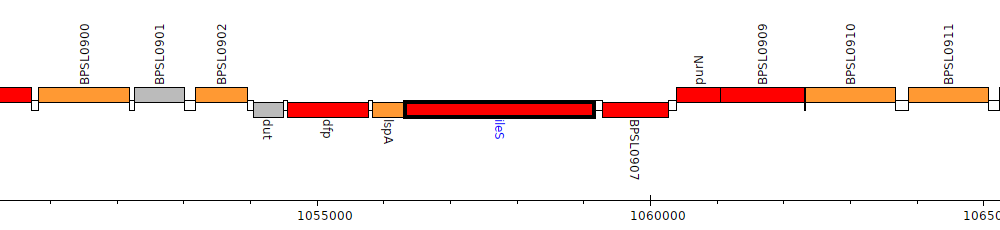
Burkholderia pseudomallei K96243, BPSL0906 (ileS)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0000049 | tRNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:cd07960
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0002161 | aminoacyl-tRNA editing activity |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.90.740.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006428 | isoleucyl-tRNA aminoacylation |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00392
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006418 | tRNA aminoacylation for protein translation |
Inferred from Sequence Model
Term mapped from: InterPro:PF00133
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000166 | nucleotide binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00133
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00133
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004812 | aminoacyl-tRNA ligase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00133
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004822 | isoleucine-tRNA ligase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00392
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bps00970 | Aminoacyl-tRNA biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00970 | Aminoacyl-tRNA biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd07960 | Anticodon_Ia_Ile_BEm | IPR033708 | Isoleucyl tRNA synthetase type 1, anticodon-binding domain | 660 | 837 | 2.06518E-80 |
| SUPERFAMILY | SSF50677 | IPR009008 | Valyl/Leucyl/Isoleucyl-tRNA synthetase, editing domain | 211 | 416 | 1.99E-47 | |
| SUPERFAMILY | SSF47323 | IPR009080 | Aminoacyl-tRNA synthetase, class Ia, anticodon-binding | 672 | 935 | 2.62E-71 | |
| PRINTS | PR00984 | Isoleucyl-tRNA synthetase signature | IPR002301 | Isoleucine-tRNA ligase | 581 | 590 | 4.7E-31 |
| Gene3D | G3DSA:1.10.730.20 | 660 | 936 | 3.6E-86 | |||
| Pfam | PF06827 | Zinc finger found in FPG and IleRS | IPR010663 | Zinc finger, FPG/IleRS-type | 907 | 932 | 1.5E-7 |
| ProSitePatterns | PS00178 | Aminoacyl-transfer RNA synthetases class-I signature. | IPR001412 | Aminoacyl-tRNA synthetase, class I, conserved site | 66 | 77 | - |
| PRINTS | PR00984 | Isoleucyl-tRNA synthetase signature | IPR002301 | Isoleucine-tRNA ligase | 59 | 70 | 4.7E-31 |
| Pfam | PF00133 | tRNA synthetases class I (I, L, M and V) | IPR002300 | Aminoacyl-tRNA synthetase, class Ia | 35 | 660 | 5.2E-187 |
| Gene3D | G3DSA:3.90.740.10 | IPR009008 | Valyl/Leucyl/Isoleucyl-tRNA synthetase, editing domain | 200 | 421 | 7.6E-217 | |
| TIGRFAM | TIGR00392 | ileS: isoleucine--tRNA ligase | IPR002301 | Isoleucine-tRNA ligase | 23 | 859 | 1.3E-259 |
| SUPERFAMILY | SSF52374 | 16 | 668 | 4.93E-126 | |||
| PRINTS | PR00984 | Isoleucyl-tRNA synthetase signature | IPR002301 | Isoleucine-tRNA ligase | 545 | 558 | 4.7E-31 |
| PRINTS | PR00984 | Isoleucyl-tRNA synthetase signature | IPR002301 | Isoleucine-tRNA ligase | 244 | 267 | 4.7E-31 |
| Hamap | MF_02002 | Isoleucine--tRNA ligase [ileS]. | IPR023585 | Isoleucine-tRNA ligase, type 1 | 10 | 936 | 20.237 |
| CDD | cd00818 | IleRS_core | 57 | 657 | 1.02691E-93 | ||
| PRINTS | PR00984 | Isoleucyl-tRNA synthetase signature | IPR002301 | Isoleucine-tRNA ligase | 414 | 429 | 4.7E-31 |
| Pfam | PF08264 | Anticodon-binding domain of tRNA | IPR013155 | Methionyl/Valyl/Leucyl/Isoleucyl-tRNA synthetase, anticodon-binding | 705 | 858 | 3.8E-25 |
| Gene3D | G3DSA:3.40.50.620 | IPR014729 | Rossmann-like alpha/beta/alpha sandwich fold | 33 | 649 | 7.6E-217 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.