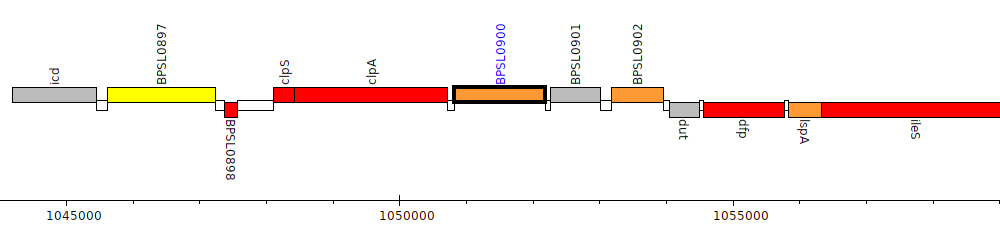
Burkholderia pseudomallei K96243, BPSL0900
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0055085 | transmembrane transport |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF006060
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016021 | integral component of membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PS00218
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006865 | amino acid transport |
Inferred from Sequence Model
Term mapped from: InterPro:PS00218
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0022857 | transmembrane transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF006060
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF006060
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PIRSF | PIRSF006060 | IPR002293 | Amino acid/polyamine transporter I | 9 | 457 | 9.3E-69 | |
| Pfam | PF00324 | Amino acid permease | IPR004841 | Amino acid permease/ SLC12A domain | 27 | 454 | 6.3E-91 |
| Gene3D | G3DSA:1.20.1740.10 | 18 | 458 | 1.1E-55 | |||
| ProSitePatterns | PS00218 | Amino acid permeases signature. | IPR004840 | Amino acid permease, conserved site | 51 | 81 | - |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.