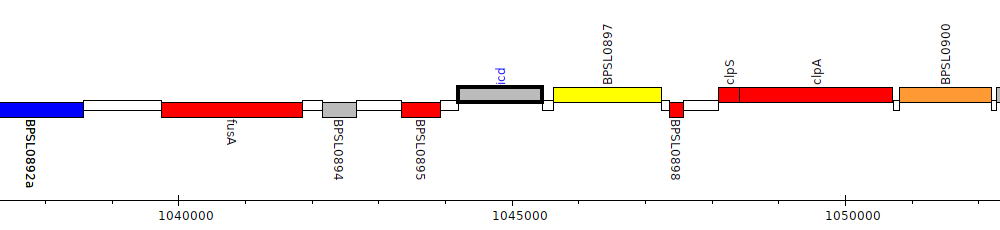
Burkholderia pseudomallei K96243, BPSL0896 (icd)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0000287 | magnesium ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS00470
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004450 | isocitrate dehydrogenase (NADP+) activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00183
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:SM01329
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006099 | tricarboxylic acid cycle |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00183
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016616 | oxidoreductase activity, acting on the CH-OH group of donors, NAD or NADP as acceptor |
Inferred from Sequence Model
Term mapped from: InterPro:SM01329
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0051287 | NAD binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS00470
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | methylaspartate cycle | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bps00020 | Citrate cycle (TCA cycle) | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | NAD/NADP-NADH/NADPH cytosolic interconversion (yeast) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | L-glutamine biosynthesis III | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bps01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | TCA cycle VII (acetate-producers) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bps01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | ethylene biosynthesis V (engineered) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bps01120 | Microbial metabolism in diverse environments | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps01230 | Biosynthesis of amino acids | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps00480 | Glutathione metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps01210 | 2-Oxocarboxylic acid metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps01200 | Carbon metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00020 | Citrate cycle (TCA cycle) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | TCA cycle V (2-oxoglutarate:ferredoxin oxidoreductase) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bps01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | partial TCA cycle (obligate autotrophs) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00480 | Glutathione metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG (InterPro) | 00720 | Carbon fixation pathways in prokaryotes | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF53659 | 4 | 418 | 8.64E-146 | |||
| Coils | Coil | 415 | 419 | - | |||
| ProSitePatterns | PS00470 | Isocitrate and isopropylmalate dehydrogenases signature. | IPR019818 | Isocitrate/isopropylmalate dehydrogenase, conserved site | 305 | 324 | - |
| Pfam | PF00180 | Isocitrate/isopropylmalate dehydrogenase | IPR024084 | Isopropylmalate dehydrogenase-like domain | 31 | 412 | 5.5E-115 |
| SMART | SM01329 | IPR024084 | Isopropylmalate dehydrogenase-like domain | 30 | 414 | 8.7E-176 | |
| Gene3D | G3DSA:3.40.718.10 | 1 | 419 | 3.0E-194 | |||
| TIGRFAM | TIGR00183 | prok_nadp_idh: isocitrate dehydrogenase, NADP-dependent | IPR004439 | Isocitrate dehydrogenase NADP-dependent, dimeric, prokaryotic | 3 | 419 | 1.3E-222 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.