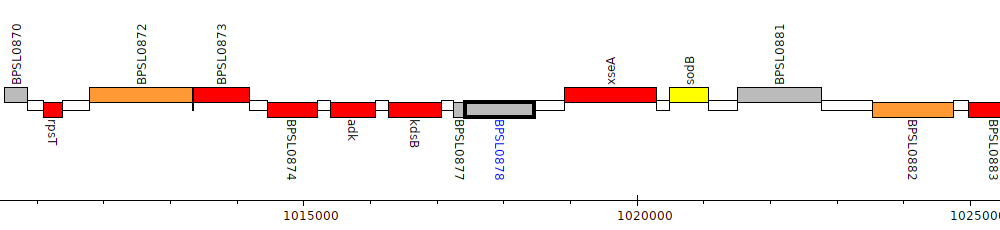
Burkholderia pseudomallei K96243, BPSL0878
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0009245 | lipid A biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00409
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00409
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0009029 | tetraacyldisaccharide 4'-kinase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00409
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG (InterPro) | 00540 | Lipopolysaccharide biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bps01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps00540 | Lipopolysaccharide biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Hamap | MF_00409 | Tetraacyldisaccharide 4'-kinase [lpxK]. | IPR003758 | Tetraacyldisaccharide 4'-kinase | 14 | 341 | 38.076 |
| Pfam | PF02606 | Tetraacyldisaccharide-1-P 4'-kinase | IPR003758 | Tetraacyldisaccharide 4'-kinase | 25 | 335 | 6.3E-99 |
| TIGRFAM | TIGR00682 | lpxK: tetraacyldisaccharide 4'-kinase | IPR003758 | Tetraacyldisaccharide 4'-kinase | 36 | 324 | 6.0E-66 |
| SUPERFAMILY | SSF52540 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 59 | 173 | 4.96E-11 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.