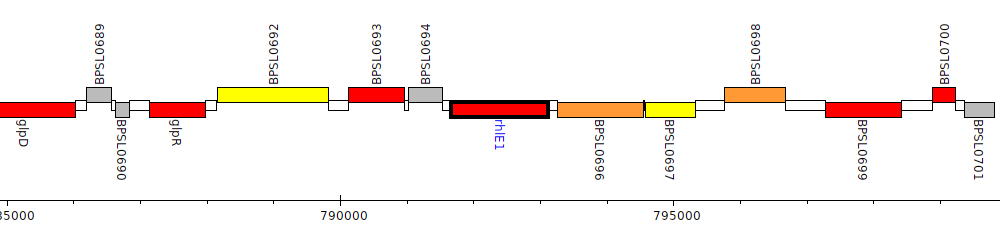
Burkholderia pseudomallei K96243, BPSL0695 (rhlE1)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003724 | RNA helicase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00968
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003676 | nucleic acid binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00270
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00270
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0042255 | ribosome assembly |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00968
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bps03018 | RNA degradation | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| ProSiteProfiles | PS51194 | Superfamilies 1 and 2 helicase C-terminal domain profile. | IPR001650 | Helicase, C-terminal | 232 | 382 | 22.579 |
| Pfam | PF00270 | DEAD/DEAH box helicase | IPR011545 | DEAD/DEAH box helicase domain | 25 | 197 | 2.3E-50 |
| Gene3D | G3DSA:3.40.50.300 | 1 | 212 | 1.7E-77 | |||
| MobiDBLite | mobidb-lite | consensus disorder prediction | 423 | 437 | - | ||
| Hamap | MF_00968 | ATP-dependent RNA helicase RhlE [rhlE]. | IPR028622 | ATP-dependent RNA helicase RhlE | 1 | 481 | 52.873 |
| SUPERFAMILY | SSF52540 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 24 | 104 | 8.5E-5 | |
| Pfam | PF00271 | Helicase conserved C-terminal domain | IPR001650 | Helicase, C-terminal | 235 | 341 | 4.6E-27 |
| Gene3D | G3DSA:3.40.50.300 | 216 | 392 | 3.4E-50 | |||
| CDD | cd00268 | DEADc | 12 | 210 | 3.19383E-102 | ||
| SMART | SM00490 | IPR001650 | Helicase, C-terminal | 260 | 341 | 2.4E-32 | |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 455 | 477 | - | ||
| ProSitePatterns | PS00039 | DEAD-box subfamily ATP-dependent helicases signature. | IPR000629 | ATP-dependent RNA helicase DEAD-box, conserved site | 155 | 163 | - |
| CDD | cd18787 | SF2_C_DEAD | 221 | 350 | 7.16809E-60 | ||
| ProSiteProfiles | PS51195 | DEAD-box RNA helicase Q motif profile. | IPR014014 | RNA helicase, DEAD-box type, Q motif | 1 | 29 | 12.56 |
| SUPERFAMILY | SSF52540 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 77 | 362 | 3.47E-75 | |
| ProSiteProfiles | PS51192 | Superfamilies 1 and 2 helicase ATP-binding type-1 domain profile. | IPR014001 | Helicase superfamily 1/2, ATP-binding domain | 32 | 209 | 31.698 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 372 | 485 | - | ||
| SMART | SM00487 | IPR014001 | Helicase superfamily 1/2, ATP-binding domain | 20 | 224 | 8.5E-60 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.