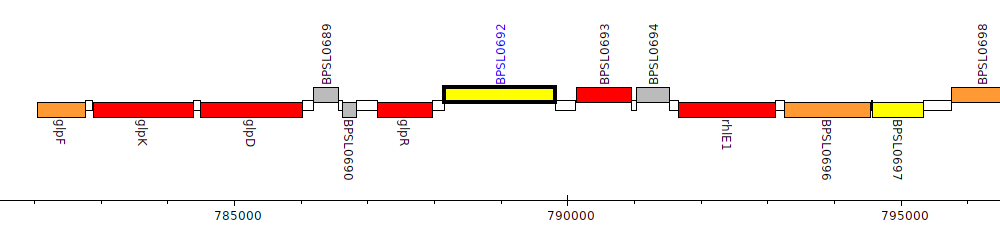
Burkholderia pseudomallei K96243, BPSL0692
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bps01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps00460 | Cyanoamino acid metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps00430 | Taurine and hypotaurine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps00480 | Glutathione metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR01210 | Gamma-glutamyltranspeptidase signature | 451 | 468 | 3.4E-44 | ||
| PRINTS | PR01210 | Gamma-glutamyltranspeptidase signature | 423 | 438 | 3.4E-44 | ||
| SUPERFAMILY | SSF56235 | IPR029055 | Nucleophile aminohydrolases, N-terminal | 17 | 557 | 7.52E-174 | |
| PRINTS | PR01210 | Gamma-glutamyltranspeptidase signature | 129 | 147 | 3.4E-44 | ||
| PRINTS | PR01210 | Gamma-glutamyltranspeptidase signature | 147 | 166 | 3.4E-44 | ||
| PRINTS | PR01210 | Gamma-glutamyltranspeptidase signature | 245 | 261 | 3.4E-44 | ||
| PRINTS | PR01210 | Gamma-glutamyltranspeptidase signature | 392 | 410 | 3.4E-44 | ||
| PRINTS | PR01210 | Gamma-glutamyltranspeptidase signature | 274 | 293 | 3.4E-44 | ||
| PRINTS | PR01210 | Gamma-glutamyltranspeptidase signature | 54 | 79 | 3.4E-44 | ||
| PRINTS | PR01210 | Gamma-glutamyltranspeptidase signature | 368 | 386 | 3.4E-44 | ||
| Gene3D | G3DSA:3.60.20.40 | 373 | 557 | 1.7E-67 | |||
| Pfam | PF01019 | Gamma-glutamyltranspeptidase | 47 | 552 | 2.8E-154 | ||
| Gene3D | G3DSA:1.10.246.130 | 261 | 372 | 2.5E-28 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.