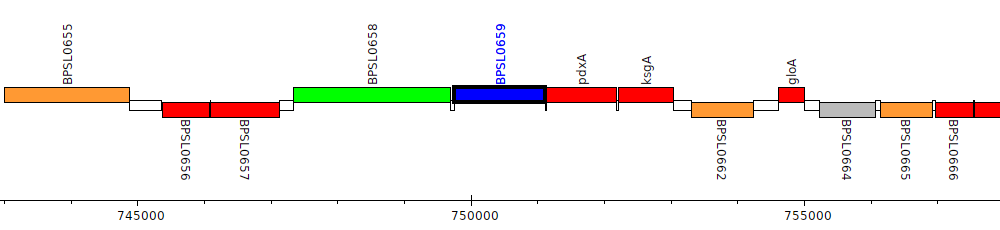
Burkholderia pseudomallei K96243, BPSL0659
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003755 | peptidyl-prolyl cis-trans isomerase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS50198
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0042277 | peptide binding |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01183
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0030288 | outer membrane-bounded periplasmic space |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01183
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0050821 | protein stabilization |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01183
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0060274 | maintenance of stationary phase |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01183
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006457 | protein folding |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01183
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0043165 | Gram-negative-bacterium-type cell outer membrane assembly |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01183
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0051082 | unfolded protein binding |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01183
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| ProSiteProfiles | PS50198 | PpiC-type peptidyl-prolyl cis-trans isomerase family profile. | IPR000297 | Peptidyl-prolyl cis-trans isomerase, PpiC-type | 303 | 401 | 26.828 |
| SUPERFAMILY | SSF54534 | 305 | 404 | 9.16E-32 | |||
| Gene3D | G3DSA:3.10.50.40 | 303 | 439 | 2.5E-39 | |||
| ProSiteProfiles | PS50198 | PpiC-type peptidyl-prolyl cis-trans isomerase family profile. | IPR000297 | Peptidyl-prolyl cis-trans isomerase, PpiC-type | 187 | 290 | 19.358 |
| Hamap | MF_01183 | Chaperone SurA [surA]. | IPR023034 | Peptidyl-prolyl isomerase SurA | 16 | 447 | 30.305 |
| Gene3D | G3DSA:3.10.50.40 | 188 | 292 | 4.8E-64 | |||
| Gene3D | G3DSA:1.10.4030.30 | 41 | 296 | 4.8E-64 | |||
| Pfam | PF00639 | PPIC-type PPIASE domain | IPR000297 | Peptidyl-prolyl cis-trans isomerase, PpiC-type | 194 | 289 | 1.4E-13 |
| Pfam | PF09312 | SurA N-terminal domain | IPR015391 | SurA N-terminal | 41 | 157 | 5.2E-32 |
| SUPERFAMILY | SSF109998 | IPR027304 | Trigger factor/SurA domain superfamily | 41 | 218 | 1.43E-40 | |
| Pfam | PF13616 | PPIC-type PPIASE domain | 302 | 401 | 1.7E-27 | ||
| SUPERFAMILY | SSF54534 | 190 | 290 | 1.11E-18 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.