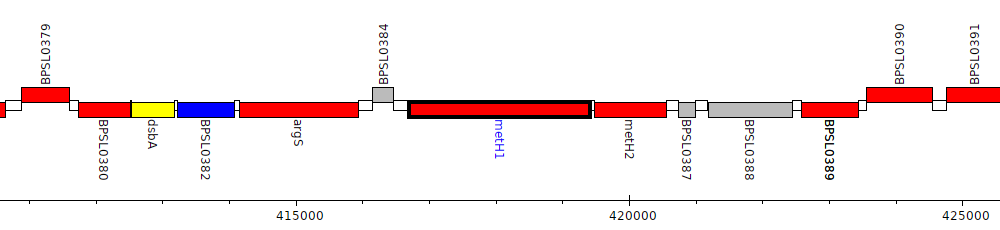
Burkholderia pseudomallei K96243, BPSL0385 (metH1)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0046872 | metal ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF02310
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0031419 | cobalamin binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF02310
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0044237 | cellular metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.20.20.20
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008705 | methionine synthase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02082
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009086 | methionine biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02082
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0042558 | pteridine-containing compound metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PS50972
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008270 | zinc ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02082
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | folate transformations I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00270 | Cysteine and methionine metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bps00670 | One carbon pool by folate | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00670 | One carbon pool by folate | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bps01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps01230 | Biosynthesis of amino acids | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00450 | Selenocompound metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | L-methionine biosynthesis IV (archaea) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bps00450 | Selenocompound metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps00270 | Cysteine and methionine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | folate transformations II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF52242 | IPR036724 | Cobalamin-binding domain superfamily | 411 | 567 | 6.93E-43 | |
| ProSiteProfiles | PS51332 | B12-binding domain profile. | IPR006158 | Cobalamin (vitamin B12)-binding domain | 415 | 554 | 42.452 |
| SMART | SM01018 | IPR003759 | Cobalamin (vitamin B12)-binding module, cap domain | 315 | 402 | 3.7E-43 | |
| ProSiteProfiles | PS50974 | AdoMet activation domain profile. | IPR004223 | Vitamin B12-dependent methionine synthase, activation domain | 569 | 900 | 83.721 |
| SUPERFAMILY | SSF51717 | IPR011005 | Dihydropteroate synthase-like | 17 | 291 | 1.23E-91 | |
| SUPERFAMILY | SSF47644 | IPR036594 | Methionine synthase domain | 310 | 401 | 3.53E-27 | |
| ProSiteProfiles | PS50972 | Pterin-binding domain profile. | IPR000489 | Pterin-binding domain | 17 | 282 | 30.943 |
| Coils | Coil | 553 | 573 | - | |||
| Pfam | PF02965 | Vitamin B12 dependent methionine synthase, activation domain | IPR004223 | Vitamin B12-dependent methionine synthase, activation domain | 610 | 885 | 4.4E-120 |
| Gene3D | G3DSA:3.40.50.280 | 412 | 568 | 2.4E-53 | |||
| SUPERFAMILY | SSF56507 | IPR037010 | Vitamin B12-dependent methionine synthase, activation domain superfamily | 575 | 900 | 9.68E-120 | |
| PIRSF | PIRSF000381 | IPR011822 | Cobalamin-dependent methionine synthase | 1 | 873 | 0.0 | |
| Gene3D | G3DSA:3.20.20.20 | IPR011005 | Dihydropteroate synthase-like | 1 | 303 | 4.8E-132 | |
| Pfam | PF02310 | B12 binding domain | IPR006158 | Cobalamin (vitamin B12)-binding domain | 417 | 510 | 1.9E-16 |
| ProSiteProfiles | PS51337 | B12-binding N-terminal domain profile. | IPR003759 | Cobalamin (vitamin B12)-binding module, cap domain | 308 | 406 | 37.674 |
| Pfam | PF02607 | B12 binding domain | IPR003759 | Cobalamin (vitamin B12)-binding module, cap domain | 317 | 394 | 2.8E-25 |
| Gene3D | G3DSA:1.10.288.10 | 609 | 771 | 3.0E-134 | |||
| Pfam | PF00809 | Pterin binding enzyme | IPR000489 | Pterin-binding domain | 21 | 259 | 4.7E-60 |
| TIGRFAM | TIGR02082 | metH: methionine synthase | IPR011822 | Cobalamin-dependent methionine synthase | 2 | 870 | 0.0 |
| CDD | cd02069 | methionine_synthase_B12_BD | IPR033706 | Methionine synthase, B12-binding domain | 317 | 544 | 2.84737E-131 |
| CDD | cd00740 | MeTr | 17 | 273 | 1.18078E-127 | ||
| Gene3D | G3DSA:3.10.196.10 | IPR037010 | Vitamin B12-dependent methionine synthase, activation domain superfamily | 580 | 900 | 3.0E-134 | |
| Gene3D | G3DSA:1.10.1240.10 | IPR036594 | Methionine synthase domain | 307 | 399 | 1.9E-38 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.