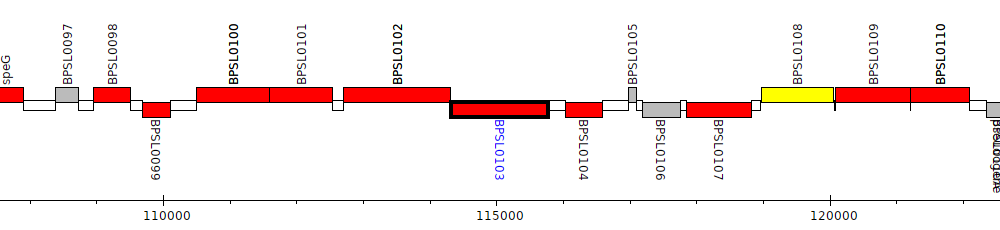
Burkholderia pseudomallei K96243, BPSL0103
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006352 | DNA-templated transcription, initiation |
Inferred from Sequence Model
Term mapped from: InterPro:PF04539
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003700 | DNA-binding transcription factor activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF04539
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006355 | regulation of transcription, DNA-templated |
Inferred from Sequence Model
Term mapped from: InterPro:PF04539
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00140
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016987 | sigma factor activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00140
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00046 | Major sigma-70 factor signature | IPR000943 | RNA polymerase sigma-70 | 454 | 465 | 9.1E-20 |
| Pfam | PF04545 | Sigma-70, region 4 | IPR007630 | RNA polymerase sigma-70 region 4 | 414 | 466 | 1.3E-14 |
| PRINTS | PR00046 | Major sigma-70 factor signature | IPR000943 | RNA polymerase sigma-70 | 294 | 302 | 9.1E-20 |
| Gene3D | G3DSA:1.10.10.10 | IPR036388 | Winged helix-like DNA-binding domain superfamily | 397 | 479 | 1.2E-17 | |
| TIGRFAM | TIGR02937 | sigma70-ECF: RNA polymerase sigma factor, sigma-70 family | IPR014284 | RNA polymerase sigma-70 like domain | 243 | 467 | 6.0E-31 |
| PRINTS | PR00046 | Major sigma-70 factor signature | IPR000943 | RNA polymerase sigma-70 | 439 | 454 | 9.1E-20 |
| PRINTS | PR00046 | Major sigma-70 factor signature | IPR000943 | RNA polymerase sigma-70 | 418 | 430 | 9.1E-20 |
| Pfam | PF00140 | Sigma-70 factor, region 1.2 | IPR009042 | RNA polymerase sigma-70 region 1.2 | 109 | 141 | 4.3E-7 |
| CDD | cd06171 | Sigma70_r4 | 409 | 466 | 7.53717E-11 | ||
| MobiDBLite | mobidb-lite | consensus disorder prediction | 41 | 80 | - | ||
| ProSitePatterns | PS00715 | Sigma-70 factors family signature 1. | IPR000943 | RNA polymerase sigma-70 | 270 | 283 | - |
| Gene3D | G3DSA:1.10.10.10 | IPR036388 | Winged helix-like DNA-binding domain superfamily | 319 | 394 | 8.8E-16 | |
| Pfam | PF04542 | Sigma-70 region 2 | IPR007627 | RNA polymerase sigma-70 region 2 | 247 | 315 | 2.0E-21 |
| ProSitePatterns | PS00716 | Sigma-70 factors family signature 2. | IPR000943 | RNA polymerase sigma-70 | 439 | 465 | - |
| Gene3D | G3DSA:1.10.601.10 | 107 | 157 | 5.1E-8 | |||
| SUPERFAMILY | SSF88946 | IPR013325 | RNA polymerase sigma factor, region 2 | 108 | 316 | 8.77E-46 | |
| Pfam | PF04539 | Sigma-70 region 3 | IPR007624 | RNA polymerase sigma-70 region 3 | 327 | 395 | 6.9E-14 |
| SUPERFAMILY | SSF88659 | IPR013324 | RNA polymerase sigma factor, region 3/4-like | 319 | 392 | 5.67E-11 | |
| SUPERFAMILY | SSF88659 | IPR013324 | RNA polymerase sigma factor, region 3/4-like | 377 | 471 | 1.31E-21 | |
| PRINTS | PR00046 | Major sigma-70 factor signature | IPR000943 | RNA polymerase sigma-70 | 270 | 283 | 9.1E-20 |
| Gene3D | G3DSA:1.10.601.10 | 179 | 318 | 3.1E-47 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.