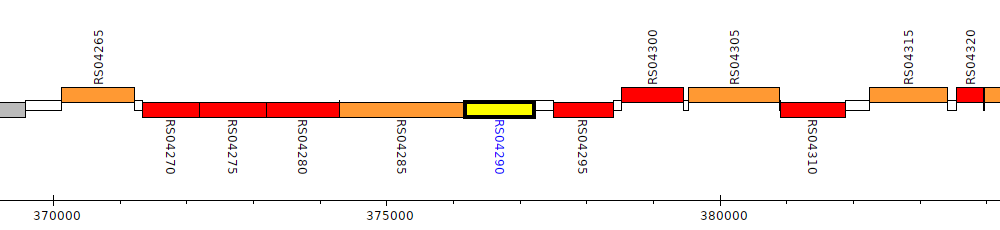
Burkholderia cenocepacia K56-2, WQ49_RS04290
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0055085 | transmembrane transport |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF006470
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0030288 | outer membrane-bounded periplasmic space |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF006470
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd13603 | PBP2_TRAP_Siap_TeaA_like | 36 | 328 | 1.78731E-84 | ||
| Gene3D | G3DSA:3.40.190.170 | IPR038404 | TRAP transporter solute receptor DctP superfamily | 24 | 323 | 4.8E-75 | |
| ProSiteProfiles | PS51318 | Twin arginine translocation (Tat) signal profile. | IPR006311 | Twin-arginine translocation pathway, signal sequence | 1 | 30 | 9.064 |
| Pfam | PF03480 | Bacterial extracellular solute-binding protein, family 7 | IPR018389 | TRAP transporter solute receptor DctP/TeaA | 56 | 317 | 7.0E-61 |
| TIGRFAM | TIGR01409 | TAT_signal_seq: Tat (twin-arginine translocation) pathway signal sequence | IPR019546 | Twin-arginine translocation pathway, signal sequence, bacterial/archaeal | 5 | 30 | 4.3E-5 |
| PIRSF | PIRSF006470 | IPR004682 | TRAP transporter solute receptor, DctP family | 6 | 332 | 7.0E-71 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.