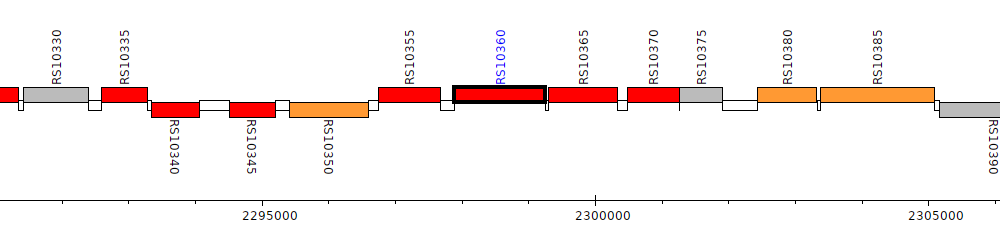
Burkholderia cenocepacia AU 1054, BCEN_RS10360
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0035596 | methylthiotransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS51449
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:SFLDS00029
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0051536 | iron-sulfur cluster binding |
Inferred from Sequence Model
Term mapped from: InterPro:SFLDS00029
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006400 | tRNA modification |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00089
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0051539 | 4 iron, 4 sulfur cluster binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00089
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016740 | transferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00089
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF01938 | TRAM domain | IPR002792 | TRAM domain | 380 | 446 | 4.0E-16 |
| ProSitePatterns | PS01278 | Methylthiotransferase radical SAM domain signature. | IPR020612 | Methylthiotransferase, conserved site | 151 | 171 | - |
| ProSiteProfiles | PS51449 | Methylthiotransferase N-terminal domain profile. | IPR013848 | Methylthiotransferase, N-terminal | 3 | 120 | 37.884 |
| TIGRFAM | TIGR01574 | miaB-methiolase: tRNA-i(6)A37 thiotransferase enzyme MiaB | 4 | 446 | 3.6E-176 | ||
| SFLD | SFLDG01082 | B12-binding domain containing | 32 | 360 | 6.5E-35 | ||
| Gene3D | G3DSA:3.40.50.12160 | IPR038135 | Methylthiotransferase, N-terminal domain superfamily | 3 | 136 | 6.4E-33 | |
| SFLD | SFLDF00273 | (dimethylallyl)adenosine tRNA methylthiotransferase (MiaB-like) | IPR006463 | tRNA-2-methylthio-N(6)-dimethylallyladenosine synthase MiaB | 2 | 446 | 0.0 |
| Pfam | PF00919 | Uncharacterized protein family UPF0004 | IPR013848 | Methylthiotransferase, N-terminal | 4 | 104 | 2.1E-29 |
| SUPERFAMILY | SSF102114 | 129 | 373 | 1.96E-60 | |||
| TIGRFAM | TIGR00089 | TIGR00089: radical SAM methylthiotransferase, MiaB/RimO family | IPR005839 | Methylthiotransferase | 4 | 443 | 3.5E-147 |
| Gene3D | G3DSA:3.80.30.20 | IPR023404 | Radical SAM, alpha/beta horseshoe | 145 | 382 | 1.8E-82 | |
| SFLD | SFLDS00029 | Radical SAM | IPR007197 | Radical SAM | 2 | 446 | 0.0 |
| ProSiteProfiles | PS50926 | TRAM domain profile. | IPR002792 | TRAM domain | 380 | 447 | 17.351 |
| Pfam | PF04055 | Radical SAM superfamily | IPR007197 | Radical SAM | 151 | 326 | 3.5E-29 |
| Hamap | MF_01864 | tRNA-2-methylthio-N(6)-dimethylallyladenosine synthase [miaB]. | IPR006463 | tRNA-2-methylthio-N(6)-dimethylallyladenosine synthase MiaB | 3 | 447 | 75.57 |
| SMART | SM00729 | IPR006638 | Elp3/MiaB/NifB | 147 | 369 | 1.2E-58 | |
| CDD | cd01335 | Radical_SAM | 156 | 358 | 3.55567E-9 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.