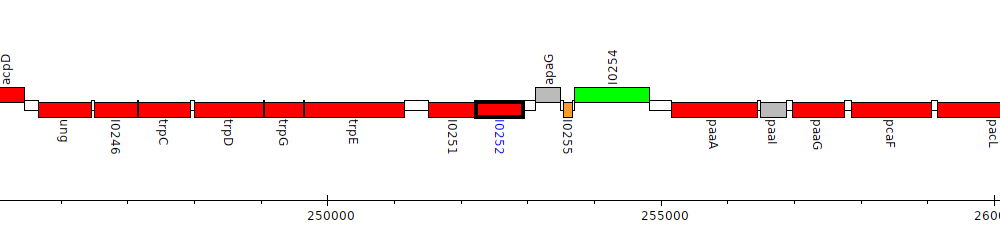
Burkholderia pseudomallei 1026b, BP1026B_I0252
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006098 | pentose-phosphate shunt |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF001461
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF51366
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016857 | racemase and epimerase activity, acting on carbohydrates and derivatives |
Inferred from Sequence Model
Term mapped from: InterPro:PS01086
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0005975 | carbohydrate metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PS01086
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004750 | ribulose-phosphate 3-epimerase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF001461
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bpz01230 | Biosynthesis of amino acids | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bpz01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bpz00040 | Pentose and glucuronate interconversions | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bpz01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bpz00710 | Carbon fixation in photosynthetic organisms | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bpz00030 | Pentose phosphate pathway | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bpz01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bpz01120 | Microbial metabolism in diverse environments | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bpz01200 | Carbon metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PIRSF | PIRSF001461 | IPR026019 | Ribulose-phosphate 3-epimerase | 1 | 229 | 1.2E-105 | |
| ProSitePatterns | PS01086 | Ribulose-phosphate 3-epimerase family signature 2. | IPR000056 | Ribulose-phosphate 3-epimerase-like | 135 | 157 | - |
| CDD | cd00429 | RPE | IPR000056 | Ribulose-phosphate 3-epimerase-like | 6 | 219 | 2.59449E-118 |
| Coils | Coil | 150 | 170 | - | |||
| TIGRFAM | TIGR01163 | rpe: ribulose-phosphate 3-epimerase | IPR000056 | Ribulose-phosphate 3-epimerase-like | 6 | 219 | 2.9E-96 |
| ProSitePatterns | PS01085 | Ribulose-phosphate 3-epimerase family signature 1. | IPR000056 | Ribulose-phosphate 3-epimerase-like | 33 | 47 | - |
| Pfam | PF00834 | Ribulose-phosphate 3 epimerase family | IPR000056 | Ribulose-phosphate 3-epimerase-like | 6 | 206 | 2.2E-90 |
| SUPERFAMILY | SSF51366 | IPR011060 | Ribulose-phosphate binding barrel | 4 | 221 | 4.1E-74 | |
| Gene3D | G3DSA:3.20.20.70 | IPR013785 | Aldolase-type TIM barrel | 1 | 226 | 3.9E-82 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.