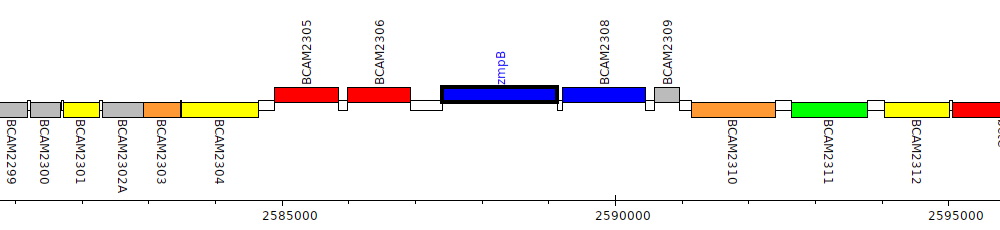
Burkholderia cenocepacia J2315, BCAM2307 (zmpB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0015628 | protein secretion by the type II secretion system | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
||
| Molecular Function | GO:0008233 | peptidase activity |
Inferred from Sequence or Structural Similarity
Term mapped from: locus_tag:BURCENK562V_RS14830
|
ECO:0000250 sequence similarity evidence used in manual assertion |
16790782 | Reviewed by curator |
| Molecular Function | GO:0008237 | metallopeptidase activity | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
||
| Molecular Function | GO:0008270 | zinc ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF02128
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004222 | metalloendopeptidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF02128
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008270 | zinc ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF02128
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0005615 | extracellular space |
Inferred from Sequence Model
Term mapped from: InterPro:PF02128
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0005615 | extracellular space |
Inferred from Sequence Model
Term mapped from: InterPro:PF02128
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004222 | metalloendopeptidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF02128
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF55486 | 284 | 558 | 3.08E-30 | |||
| Pfam | PF02128 | Fungalysin metallopeptidase (M36) | IPR001842 | Peptidase M36, fungalysin | 397 | 520 | 2.7E-6 |
| Pfam | PF07504 | Fungalysin/Thermolysin Propeptide Motif | IPR011096 | FTP domain | 107 | 153 | 4.7E-10 |
| Gene3D | G3DSA:3.10.450.490 | 63 | 217 | 3.2E-5 | |||
| Gene3D | G3DSA:3.10.170.10 | 258 | 416 | 8.1E-5 | |||
| CDD | cd09596 | M36 | 397 | 497 | 3.59591E-5 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.