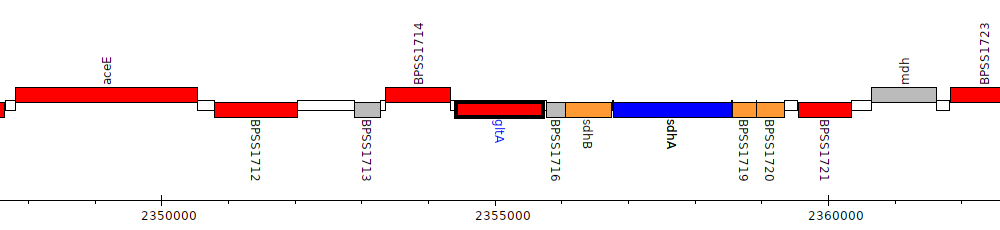
Burkholderia pseudomallei K96243, BPSS1715 (gltA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0046912 | transferase activity, transferring acyl groups, acyl groups converted into alkyl on transfer |
Inferred from Sequence Model
Term mapped from: InterPro:PF00285
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006099 | tricarboxylic acid cycle |
Inferred from Sequence Model
Term mapped from: InterPro:PS00480
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004108 | citrate (Si)-synthase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01798
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0005737 | cytoplasm |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01798
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bps01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps00630 | Glyoxylate and dicarboxylate metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps01200 | Carbon metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps00020 | Citrate cycle (TCA cycle) | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps01230 | Biosynthesis of amino acids | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps01210 | 2-Oxocarboxylic acid metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps01120 | Microbial metabolism in diverse environments | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PIRSF | PIRSF001369 | IPR024176 | Citrate synthase, bacterial-type | 41 | 433 | 8.2E-169 | |
| PRINTS | PR00143 | Citrate synthase signature | IPR002020 | Citrate synthase | 364 | 380 | 5.2E-55 |
| Gene3D | G3DSA:2.20.28.60 | 5 | 54 | 8.6E-26 | |||
| PRINTS | PR00143 | Citrate synthase signature | IPR002020 | Citrate synthase | 300 | 320 | 5.2E-55 |
| ProSitePatterns | PS00480 | Citrate synthase signature. | IPR019810 | Citrate synthase active site | 307 | 319 | - |
| Gene3D | G3DSA:1.10.230.10 | IPR016143 | Citrate synthase-like, small alpha subdomain | 272 | 381 | 1.5E-147 | |
| PRINTS | PR00143 | Citrate synthase signature | IPR002020 | Citrate synthase | 245 | 273 | 5.2E-55 |
| PRINTS | PR00143 | Citrate synthase signature | IPR002020 | Citrate synthase | 384 | 398 | 5.2E-55 |
| TIGRFAM | TIGR01798 | cit_synth_I: citrate (Si)-synthase | IPR010953 | Citrate synthase, type I | 18 | 425 | 4.9E-242 |
| Gene3D | G3DSA:1.10.580.10 | IPR016142 | Citrate synthase-like, large alpha subdomain | 61 | 416 | 1.5E-147 | |
| PRINTS | PR00143 | Citrate synthase signature | IPR002020 | Citrate synthase | 223 | 238 | 5.2E-55 |
| PRINTS | PR00143 | Citrate synthase signature | IPR002020 | Citrate synthase | 169 | 182 | 5.2E-55 |
| SUPERFAMILY | SSF48256 | IPR036969 | Citrate synthase superfamily | 9 | 433 | 2.49E-169 | |
| Pfam | PF00285 | Citrate synthase, C-terminal domain | IPR002020 | Citrate synthase | 50 | 416 | 4.9E-134 |
| CDD | cd06114 | EcCS_like | 21 | 421 | 0.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.